Contact Us
Latest News
- Clues beginning to emerge on asymtomatic SARS-CoV-2 infection
- Back in November of 2020, during the first wave of the COVID-19 pandemic, I was teaching an in-person microbiology laboratory. One of my students had just been home to see his parents, and they all c…
- Read more
- Could there maybe be better uses of genetics and probiotics?
- Professor Meng Dong and his laboratory have created a probiotic that can metabolize alcohol quickly and maybe prevent some of the adverse effects of alcohol consumption. The scientists cloned a highl…
- Read more
- ChatGPT is not the end of essays in education
- The takeover of AI is upon us! AI can now take all our jobs, is the click-bait premise you hear from the news. While I cannot predict the future, I am dubious that AI will play such a dubious role in…
- Read more
- Fighting infections with infections
- Multi-drug-resistant bacterial infections are becoming more of an issue, with 1.2 million people dying of previously treatable bacterial infections. Scientists are frantically searching for new metho…
- Read more
- A tale of two colleges
- COVID-19 at the University of Wisconsin this fall has been pretty much a non-issue. While we are wearing masks, full in-person teaching is happening on campus. Bars, restaurants, and all other busine…
- Read more
Chapter 3 - Cell Structure and Organization
3 - 1 Introduction
Microorganisms typically face the world as single cells rather than the multi-cellular assemblies of higher organisms. Therefore, each cell must contain all the structures necessary for managing its internal state and dealing with the outside environment. Not surprisingly, this evolutionary process results in similar structures and processes to solve similar needs in different microorganisms. However, prokaryotes have been on this earth for a long time, and this has allowed them to differentiate into a dizzying number of different species. Eukaryotic microbes are not quite so diverse, but they still display a remarkable range of properties. No one cell contains all the structures that we describe here, but we will explore the more common structures (Figure 3.1) observed by scientists in the past 150 years. A distinction in this discussion must be made between the two types of prokaryotes: the Archaea and their cousins, the Bacteria. We will initially focus on Bacteria since we know the most about them. Many of the structures we will examine are present in Bacteria and Archaea, but there are significant differences. We will cover these at the end of the chapter. Finally, we will talk about the features that are distinctive among the microbial eukaryotes.
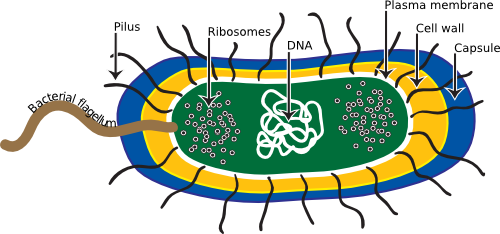
Figure 3.1. The Generalized Bacteria. This cartoon displays many of the common structure found in prokaryotic microorganisms, though not every one will have every one of these structures.
So how did scientists find out so much about such tiny organisms? As you might guess, many techniques come into play when tackling a subject as complex as bacterial structure. Microscopes help scientists visualize the location of structures and how they are arranged spatially in the microbe. Bacterial genetics and molecular biology identify and analyze the genes necessary for the synthesis and regulation of these structures. Biochemistry permits the detailed examination of each part separately, with implications for its role in the living bacterium. The powerful combination of these disciplines provides a deep understanding of how a bacterium is put together, but there is still much to learn.
3 - 2 Important structures in the cytoplasm
Learning Objectives
After reading this section, students will be able to...
- Explain the organizational units of the cell, cytoplasm, ribosomes and nuclear region.
- Describe how enzymes in a pathway can be organized into structures that facilitate their activity.
- Identify the structure of the cytoplasm and explain that most of the metabolic reactions occur in the cytoplasm.
- Explain the roles of RNA in translation.
- Explain the roles of microscopically visible structures in the cell (inclusions, vesicles, and magnetosomes).
For a refresher or introduction to basic chemistry and biochemistry, please read chapter 2. In this chapter, we will look at how this chemistry combines to form major functional units. These units carry out the major business of the cell: growth, replication, feeding, and movement. We will describe the cell from the inside out, starting with the cytoplasm, moving to the membrane, the cell wall, and finally surface structures on microbes. Before we begin, we should describe an important evolutionary hypothesis that will make sense of much of the following details. As you will see, there are a number of curious similarities and differences in the details of cellular structure among Bacteria, Archaea and eukaryotes. In general much of the machinery in a eukaryotic nucleus and in the cytoplasm looks rather a lot like what is present in the archaea. However, the organelles of eukaryotes, such as the mitochondria and chloroplasts, have properties that are much more similar to those of bacteria. How is this possible? One clue comes from observing organisms in nature. Cooperative relationships are ubiquitous between different species, and this is also true in the microbial world. In some instances, these relationships involved close physical contact between their participants, sometimes with one participant engulfing the other. In 1968 Dr. Lynn Margulis extended this observation and proposed that some of the organelles found in eukaryotes, specifically mitochondria and chloroplasts, were originally endosymbionts of their host. Originally these two microbes probably could live independently, but over time, the endosymbiont lost functionality that its host was already providing and then became dependent. Over the years, ample evidence has accumulated to support this exceptional insight.
- Both mitochondria and chloroplasts contain DNA that resembles the chromosomes of bacteria.
- Both organelles are surrounded by two membranes reminiscent of Gram-negative cell wall structure (see below).
- Mitochondria and chloroplasts divide by a method that resembles binary fission.
- Much of the internal structure and biochemistry of the photosynthetic organelle inside chloroplasts is very similar to that observed in cyanobacteria (a photosynthetic microbe).
- The ribosomes of mitochondria and chloroplasts resemble those found in microbes and analysis of the sequence of the 16S rRNA of these ribosomes showed that the organelles are in fact closely related to proteobacteria (mitochondria) and cyanobacteria (chloroplasts) This last bit of proof is very strong evidence substantiating Dr. Margulis's hypothesis.
In summary, it seems clear that the eukaryotic cell was born by merging an archaeal cell with a Gram-negative proteobacteria. Photosynthetic eukaryotes arose from a second endosymbiosis, where the eukaryotic cell engulfed a cyanobacterium. Clearly, eukaryotes, including us, resulted from the cooperation of several single-celled species in the long distant past. In other words, not only do you have bacteria growing on you, you are bacteria. Therefore, what you learn here about the basic workings of a bacterial cell will apply to all cells.
The cytoplasm is the area inside the membrane
The cytoplasm or protoplasm is the portion of the cell that lies within the cytoplasmic membrane. The cytoplasmic matrix consists of substances within this membrane, excluding the genetic material. In most prokaryotes, it appears to be relatively featureless by electron microscope, but all that means is that there are no large structures we can see. In contrast, eukaryotic cells have mitochondria and other visible organelles that exist for different specific functions. Despite this visual simplicity, the prokaryotic cytoplasm is the site of almost all important metabolic functions in the cell.
The cytoplasm has a gel-like consistency, with rather different properties than the simple solutions that we typically make up in the laboratory. This characteristic is because there is surprisingly little free water in the cell. Rather than picturing the cytoplasm as a pool of water with the occasional large molecule floating around, it is better to think of it as a bag of proteins and other macromolecules, each coated with a layer of water, with a modest number of free water molecules bouncing around in between. There is so little free water in the cell that one-third of all water molecules are making hydrophilic contact with the macromolecules in the cell. Given this difference between our lab solutions and the actual nature of the cytoplasm, it is a bit surprising that the biochemical analyses we perform in the lab mimic the behavior observed in the cell.
Enzymes serve as catalysts in the cytoplasm
The cytoplasm is where most metabolism occurs, and we describe the details of these processes in the chapters on metabolism and photosynthesis. However, we mention few general issues concerning enzymes in the cytoplasm here.
Though the cytoplasm appears featureless, there is a significant amount of local organization. A good illustration of this is exemplified by examining the enzymes of DNA replication. Though too small to be seen, the proteins that perform replication are in complex assemblies of many proteins. This assembly is much more efficient than having each protein float around and simply sump into the DNA to perform its function by random chance. For example, DNA gyrase, which unwinds and opens the DNA for copying, has to function in coordination with DNA polymerase, which inserts each new nucleotide in the growing strand. Without this coordination, the DNA would not open up for replication, and the process would simply not occur. It would certainly be possible for the enzymes to float around in the cytoplasm without any interaction in some other processes. Still, it is much more efficient to organize in some fashion. The glycolytic enzymes (enzymes that oxidize sugars for energy) exemplify this type of multi-enzyme complex. One enzyme directly hands over its product to the next enzyme, for which it is the substrate - a sort of molecular assembly line. This coordination of functions is much more efficient because substrate does not accumulate where it should not, and the local substrate concentration for each enzyme is very high.
The cell DNA is organized into a nucleoid
The nucleoid is the cellular region that contains the DNA-protein complex that makes up the chromosome. It is not a set region, as in eukaryotes, with their nuclei but can vary from cell to cell. So what do we know about prokaryotic chromosomes? What is the structure of a typical chromosome?
It is a little awkward to talk about "typical" anything in prokaryotes, since there is always such a large range for any given property. In the case of prokaryotic chromosomes, the smallest known is about 160 kB, while the largest is about 10,000 kB. E. coli is certainly well-studied and might well be considered typical, with strain K12 having a chromosome of 4,639,221 bp containing 4,405 genes. The chromosome for this bacterium is circular, and this is a common arrangement, but there are many species with linear chromosomes. If stretched out, this material would be about 1400 µm long or about 1/16thof an inch. The E. coli cell is about 1-5 µm in length, so it is clear that a remarkable degree of packing is necessary to fit the chromosome inside its tiny host. Figure 3.2 shows the nucleoid of E. coli. The nucleoid occupies half of the bacterial cytoplasm and has a density of 20-50 mg/ml, similar to what is observed for the nucleus of a non-dividing eukaryotic cell. The following description of the nucleoid refers specifically to that of E. coli, but is probably generally similar to that of other prokaryotes.
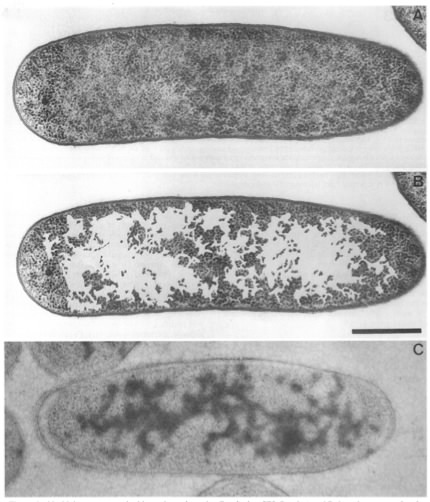
Figure 3.2. Electron micrograph of the nucleoid. The nucleoid as seen in thin sections of growing E. coli. Panels A and B show the same section; in panel B the ribosome-free spaces were enhanced by coloring by hand. Panel C shows a similar cell stained with antibodies specific for DNA. (Source: E. Kellenberger).
The nucleoid is composed of DNA associated with several DNA-binding proteins that help it maintain its structure. The proteins HU, H-NS, and IHF are found in the DNA and help create the nucleoid's structure. HU and H-NS non-specifically bind to DNA, with H-NS serving as the major DNA binding protein. IHF and HU both facilitate the bending of DNA, but IHF does so by binding to specific DNA sites. The nucleoid also contains a large amount of RNA polymerase and RNA, as well as small amounts of many different proteins that regulate the expression of specific genes. These seem not to perform any structural role but reflect the importance of RNA transcription in the nucleoid in growing cells.
The DNA double helix typically has a bit of twisting tension in the opposite direction of the helix ladder (the twists typically "unwind" the left-handed DNA helix). Negative supercoiling, as it is called, is produced by the action of enzymes termed topoisomerases. Figure 3.3 shows the supercoiling of the DNA. This negative supercoiling makes it slightly easier to separate the two strands of the double helix, as must be done to start transcription and replication.
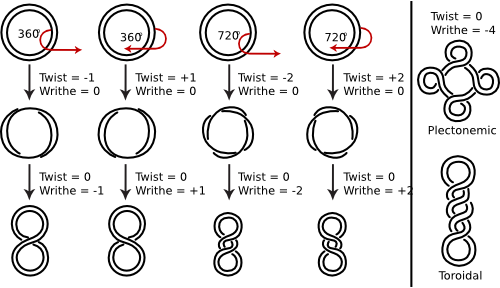
Figure 3.3. Supercoiling of DNA strands. Supercoiling is very difficult to show in a static picture and it is hard to create a cartoon that adequately describes it. Instead consider this: Take a few feet of tubing or a garden hose and hold one end in your left hand. Then with your right hand, rotate the other end along its long axis. Since the tubing/hose is resistant to this twisting, this energy will cause a section of the tubing to form a new structure, a bit like the middle picture above. This "supertwisting" is supercoiling. Now the analogy between tubing and a DNA double-helix is not bad - both objects are able to bend but cannot really rotate much along their long axis, so if one could do the same thing with a piece of DNA, the same supertwisting would result. Of course no one is "holding one end of the DNA" in the cell, but the fact that the DNA is a closed circle has the same effect. In this picture, double-stranded DNA is shown, but for clarity the helical nature of the DNA is not depicted (though it certainly IS helical). Twist refers to the number of helical turns in the DNA and writhe to the number of times the double helix crosses over on itself (these are the supercoils). The bottom two figures shows how the supercoils could be concentrated in a single region or dispersed.
Because supercoiling changes fairly rapidly, studying it in living cells has been challenging. As shown in Figure 3.4,the chromosome is further folded into 50 or so loops of about 100,000 base pairs. These domains supercoil independently, and indeed, even small sections within the same loop can transiently have different degrees of supercoiling. It does appear that supercoiling can affect the expression of genes in a region of the DNA and that gene expression also affects supercoiling. In addition to this rosette morphology, one or several loci of the chromosome are specifically positioned within the cell, and the intervening DNA is kept organized with respect to these landmarks. In fact, you will find specific regions of the chromosome in similar positions in the cell. How this is specifically arranged is species-specific, with different microbes arranging their chromosomes in different ways. For example, E. colikeeps its oriCat the center of the cell and has left and right halves of the chromosome that flow outward from there, eventually meeting at the terminus. Caulobacterspecies, curved rod-shaped microbes, will keep their origin and terminus at opposite poles of the cell.
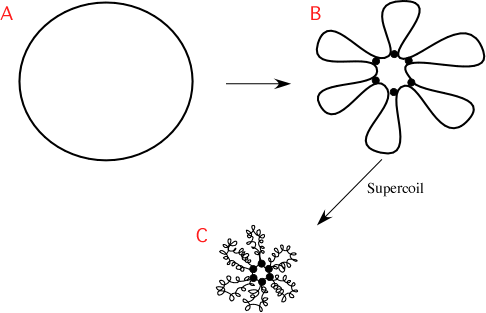
Figure 3.4. Chromosome structural organization. A model of the overall structure of the bacterial chromosome. (A) The unfolded, circular chromosome of E. colidepicted as a single line for simplicity, though of course it is a double-stranded helix. (B) The DNA folded into chromosomal domains by protein-DNA associations. The proteins are depicted as the black circles, interacting with both the DNA and with each other. Six domains are shown, but the actual number for E. coli is about 50. (C) Supercoiling and other interactions cause the chromosome to compact greatly.
Transcription and translation occur on the surface of the nucleoid
So we have told you that the DNA in the cell has a very compact structure because of DNA-binding proteins, but we have also said that this structure is very dynamic because gene transcription is going on all of the time. This paradox creates a complicated situation for the cell that only worsens when it is time to replicate the chromosome. Remember that prokaryotes continue to perform gene expression throughout replication, in contrast to eukaryotes. As you will learn below, transcription and translation are coupled in bacteria - the beginning of the messenger RNA (mRNA), termed the 5' end, is being actively translated while the last portion, termed the 3' end, is still being synthesized (Figure 3.5). Yet, the internal regions of the nucleoid appear to be devoid of ribosomes and non-DNA-binding proteins, suggesting that all transcription and translation must occur on the surface of the nucleoid. Therefore the cell must shift this large tangle of DNA around by some unknown mechanism as gene expression of certain buried sequences is needed. It is a marvel that the whole process of transcription, translation, and replication works at all, especially within the tiny confines of the cell.
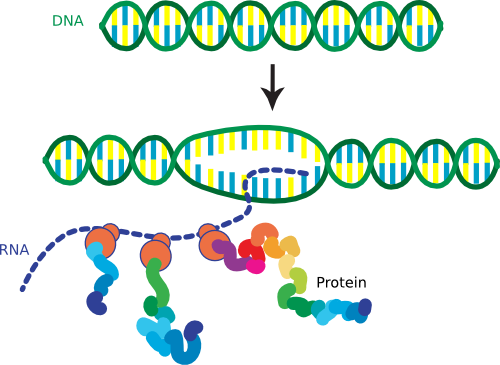
Figure 3.5. The coupling of transcription and translation. In prokaryotes the process of transcription and translation are tightly coupled. This increases the rate at which proteins can be expressed and is one reason that some bacteria can multiply so quickly.
Translation involves messenger RNA
Translation is the process of converting the instructions coded in the DNA into the proteins that carry out the work. The macromolecules that perform this task consist of mRNA, transfer RNA (tRNA), and the ribosome, composed of ribosomal RNA (rRNA) and ribosomal proteins. In brief, this process consists of making an mRNA copy of a region in the DNA that gives directions for the synthesis of protein or proteins. The ribosome then binds the mRNA and translates it into an amino acid sequence. The amino acids necessary for the protein are carried to the ribosome by tRNA that read the information in the mRNA and add the appropriate amino acid to the nascent protein chain. This process is described in much greater detail in the chapter on the central dogma.
We will now examine the structure of the molecules involved in translation, starting with mRNA, where the primary structure is simple - merely unmodified A, G, C and U bases. In almost all prokaryotic mRNAs there is not a great deal of secondary and tertiary structure, since ribosomes are typically translating them and the translating ribosome removes any structure as it moves along the mRNA. What structure there is occurs in the untranslated regions, notably the 5' and 3' ends of the mRNA. One of these structural roles, especially at the 3' end, is to stabilize the mRNA. There are RNases in the cell that degrade the RNA and RNA secondary structure can impede this degradation. The amount of secondary structure is by design, depending upon how long the cell wants the mRNA to last. Due to the need of most prokaryotic cells to respond rapidly to changes in the environment, most mRNAs are not very stable in the cell and degrade rapidly.
Ribosomes are composed of RNA and protein
In contrast to the case with mRNAs, the other RNAs involved in translation, tRNA and rRNA, have very distinct structures. Each rRNA folds into a known secondary structure and has a complex tertiary structure containing many short helical regions and long-range base pair interactions. Interactions between the RNAs and proteins also maintain these structures.
The composition of ribosomes is 62 % RNA and 38 % protein by weight. Two complexes of RNA and protein make up the ribosome, the 30S subunit, and the 50S subunit. (The S stands for Svedberg units, a measure of how fast something sediments in solution. All you need to know for our purposes is that larger molecules sediment faster and have larger Svedberg units. The unit is named after Theodor Svedberg, who won the Nobel Prize in Chemistry in 1926 for his work on suspensions of large molecules and other compounds in solutions.) The 30S subunit is composed of 21 proteins and a single-stranded rRNA molecule of about 1,500 nucleotides, termed the 16S rRNA. The 50S subunit contains 31 proteins and two RNA species, a 5S rRNA of 150 nucleotides and a 23S rRNA of about 2,900 nucleotides. Several of the nucleotides on the 16S and 23S rRNAs have been modified by methylation. These modifications are probably critical to the function of the rRNAs since they always happen in regions conserved through evolution.
The ribosome-associated proteins are positively charged, with a high proportion of lysine and arginine residues. This positive charge facilitates complex formation between the acidic RNA and these basic proteins. Scientists have solved the crystal structure of the entire 70S ribosome, as shown in Figure 3.6. It is remarkable because solving such structures becomes more difficult as the size of the complex increases and the ribosome is huge. We are still trying to understand what the structure tells us about ribosome function. We also know that the ribosome is not static but dynamic, changing shape during a translation cycle.
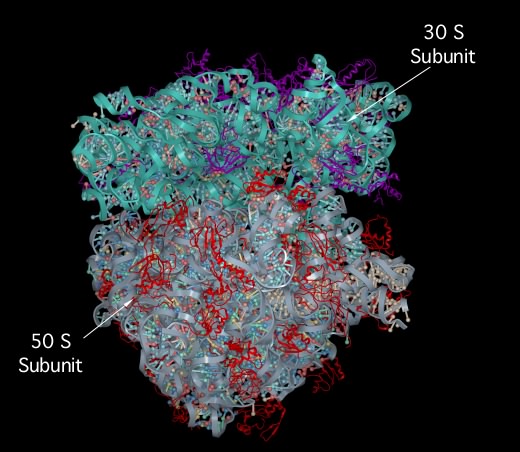
Figure 3.6. The crystal structure of the 70S ribosome.. A molecular model of the 70S ribosome of Thermus thermophilis showing the position of proteins (purple and red ribbon diagrams) and RNA (Gray and cyan ribbons) within the structure. Note the prominence of RNA in the ribosome, which constitutes over 60% of the molecular weight. (Source: adaption from S. Petry et al. 2005. Cell 123:1255-1266)
Transfer RNA is the ferry for amino acids
Transfer RNA (tRNA) is the ferry that transports the amino acids to the ribosome. There are one or more different tRNA molecules for each of the 20 amino acids. Each consists of 70 to 80 nucleotides of single-stranded RNA that is extensively base-paired to form four short helical domains. These structures are commonly represented as a two-dimensional cloverleaf but look more like an "L" in the native three-dimensional structure, as shown in Figure 3.7. The tertiary structures of tRNAs are similar, so the critical features that make each appropriate to a specific amino acid are largely found in the primary structure itself. Enzymes chemically modify many bases in tRNA molecules to help the molecule carry out its function.
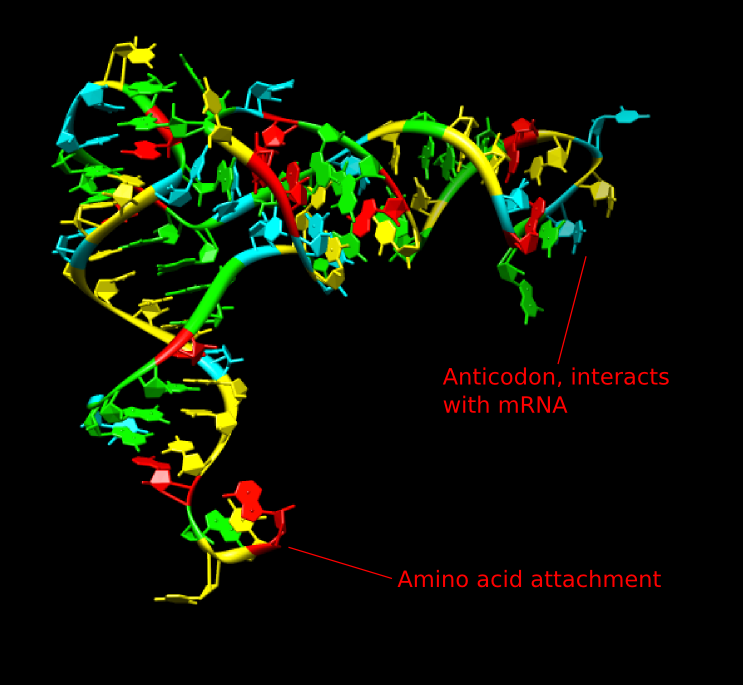
Figure 3.7. The structure of tRNA. The two-dimensional structure of tRNA looks like a cloverleaf, but in the actual three-dimensional form, it has a surprising L-shaped structure as shown. The bottom of the structure as shown here contains the anticodon that interacts with the mRNA. The top right of the structure is where the appropriate amino acid residue is attached by synthetases.
The aminoacyl-tRNA synthetases are the enzymes that add the amino acid to the tRNAs. Figure 3.8 depicts a complex between an aminoacyl tRNA synthetase and tRNA. There is a single synthetase for each amino acid, and it binds each of its appropriate tRNA molecules and charges it with its appropriate amino acid. The synthetases avoid both the wrong tRNAs as well as the wrong amino acids.
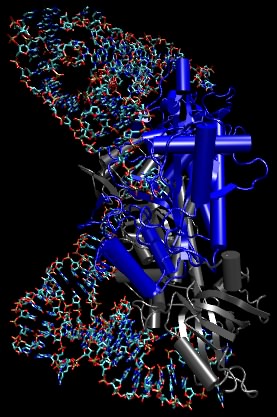
Figure 3.8. The amino acyl-tRNA synthetase complex. A molecular model of aminoacyl tRNA synthetase binding its tRNA. The tRNA is shown as ball-and-stick, while the synthetase is depicted in the ribbon form. In this picture, the anticodon is at the top of the figure and the site of amino acid attachment is at the bottom right. Note that the synthetase does not "sense" the anticodon directly.
One final thought before we move on. Think about the chicken-and-egg conundrum that translation brings up. This whole process has the express purpose of synthesizing proteins, yet it involves proteins at every step. How could these proteins have evolved to serve this function when they are necessary for their synthesis? In other words, how could you synthesize anyprotein until a complete set of translation proteins had evolved? Part of the answer is that primordial translation was probably much simpler, though less accurate and efficient. Perhaps only very few and somewhat general protein functions were required. Alternatively, perhaps early translation used no proteins at all: Some scientists believe that early life employed RNA molecules capable of carrying out necessary enzymatic functions and storing hereditary information. They posit that it was only later that proteins came along and assisted in their synthesis. However, this hypothesis still does not explain how one simultaneously evolved functional proteins and a process for creating them.
Inclusions and other internal structures are found in many prokaryotic cells
There are a few structures in the prokaryotic cytoplasm that are visible by microscopy. Generally, they serve specific purposes in the cell and are often found only in specific cell types or under certain growth conditions. Here we focus on some of the more familiar objects, inclusions, gas vesicles, and magnetosomes.
Inclusions
Inclusions are dense aggregates of specific chemical compounds in the cell. Typically, the aggregated chemical serves as a reservoir of either energy-rich compounds or building blocks for the cell. Forming polymers costs energy, and it may seem wiser for the cell to keep the excess monomers around for when they are needed. The benefit of polymerization is that it decreases the osmotic pressure on the cell, a serious problem as described later. Inclusions often accumulate under laboratory conditions when a cell is grown in the presence of excess nutrients. However, the role of some inclusions is unclear. Growth on rich medium causes their creation, but subsequent starvation in the test tube does not always result in the use of these reserves. This lack of use suggests that these inclusions, at least, are not storage bodies or that we have not found conditions where the cell will use them.
Carbon and energy storage
One of the more common storage inclusions involves poly-β-hydroxyalkanoate(PHA). It is a long polymer of repeating hydrophobic units that can have various carbon chains attached to it. The most common form of this class of polymers is poly-β-hydroxybutyrate, which has a methyl group as the molecule's side chain, as shown in Figure 3.9. The function of PHA in bacteria is as a carbon and energy storage product. Just as we store fat, some bacteria store PHA. Some PHA polymers have plastic-like qualities, and there is some interest in exploiting them as a form of biodegradable plastic.
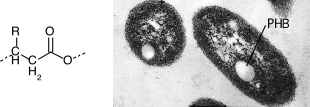
Figure 3.9. Poly-β-hydroxyalkanoate inclusion bodies. The figure shows an electron micrograph of inclusion bodies of poly-β-hydroxyalkanoate inside a cell of a Rhodobacter sphaeroides. The specific chemical here is poly-β-hydroxybutyrate (PHB). In the generalized poly-β-hydroxyalkanoate structure shown at the left, the R group is a methyl in PHB. (Source, Sam Kaplan, University of Texas - Houston Medical School)
Glycogen is another common carbon and energy storage product. Humans also synthesize and utilize glycogen, which is a polymer of repeating glucose units.
Phosphate granules and sulfur globules
Given the opportunity, many organisms accumulate granules containing long phosphate chains since this is often a limiting nutrient in the environment. These polyphosphate polymers, also called volutin, form visible granules in some microbes. These granules are readily stained by many basic dyes such as toluidine blue and turn reddish violet in color. These inclusions are often called metachromatic granules because they become visible by "metachromasy" (a color change). Polyphosphate is found in all known cells (eukaryotes, bacteria and archaea) and appears to serve many important roles.
- It serves as a phosphate reservoir
- It is an alternative substrate in place of ATP when phosphorylating sugars during catabolism.
- It is a chelator for divalent cations
- It can be a buffer under alkaline stress
- It is an important factor for DNA uptake.
- Finally, phosphate polymers are important regulators in response to stress
Figure 3.10 depicts another visible structure, termed a sulfur globule, which is present in various bacteria capable of oxidizing reduced sulfur compounds such as hydrogen sulfide and thiosulfate. Oxidation of these compounds is linked either to energy metabolism or photosynthesis. Oxidation of sulfide initially yields elemental sulfur, which accumulates in globules inside or outside the cell. If the bacterium exhausts the sulfide in the area, it may further oxidize the sulfur to sulfate.
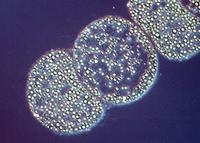
Figure 3.10. Sulfur globules. Sulfur globules found in Thiomargarita namibiensis. This large microbe (100 to 750 µm in size) is found in Walvis Bay off the coast of Namibia. (Source: copyright Max Planck Institute for Marine Microbiology, Bremen, Germany)
Gas vesicles
Figure 3.11 shows an example of gas vesicles, also known as gas vacuoles, that are found in cyanobacteria. Cyanobacteria are photosynthetic and live in lakes and oceans. In these environments, the cyanobacteria use gas vesicles to control their position in the water column to obtain the optimum amount of light and nutrients.
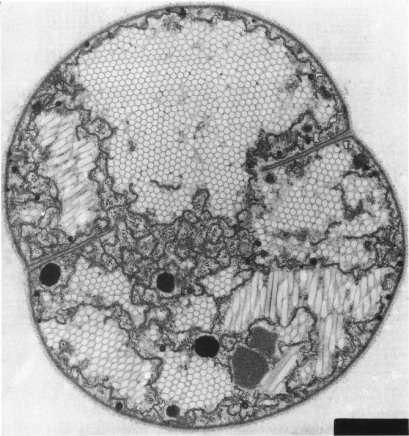
Figure 3.11. Gas vesicles. The hexagonal forms inside the cytoplasm of this cyanobacterium are the gas vesicles. These actively dividing cells are Microscystissp.(Source: A. E. Walsby, 1994. Microbiol. Rev. 58:94-144)
Gas vesicles are often aggregates of hollow cylindrical structures composed of rigid proteins. They are impermeable to water, but permeable to gas. The amount of gas in the vesicle is under the control of the microorganism. The release of gas from the cell causes the bacteria to fall in the water column while filling the vesicle with gas causes the cells to rise.
Magnetosomes
Magnetosomes are intracellular crystals of iron magnetite (Fe3O4) that impart a permanent magnetic dipole to prokaryotic cells that have them. They allow these microbes to orient themselves in a magnetic field. This process does not appear to involve any special machinery besides the magnetosome. Each microbe can be thought of as having a tiny magnet that responds to the magnetic field in the environment. These magnetosomes allow the microbes to follow the magnetic field of the earth. Some species of magnetotactic bacteria have the following behavior. In the northern hemisphere, magnetotactic bacteria swim north along the magnetic field, while in the southern hemisphere, they swim south. Because of the inclination of the earth's magnetic field, this causes the microbes to swim downward. Many microbes containing magnetosomes are aquatic organisms that do not grow well in atmospheric concentrations of oxygen. They move away from the oxygen higher up in the water column by detecting the magnetic field and swimming downward.
A special membrane surrounds magnetosomes that confine the magnetite to a defined area. The membrane likely plays a role in precipitating the iron as Fe3O4 in the developing magnetosome. Magnetosomes can be square, rectangular or even spike-shaped.
Key Takeaways
- The cytoplasm is that area inside the membrane and has a gel-like consistency.
- While there are no visible structures in the cytoplasm in prokaryotes, it is still highly organized. As an example, the nucleoid of the cell, containing the chromosome, is bound by proteins and supercoiled to tightly pack the DNA into the cell.
- Translation occurs at the ribosome and converts the recipe of the protein, encoded in DNA, and mRNA into protein.
- This process also involves another RNA, tRNA, as an adapter.
- The cell cytoplasm does contains some visible structures, called inclusions. These structures function as nutrient storage (PHA, sulfur and phosphorus globules), buoyancy (gas vesicles), and sensing the earth's magnetic field (magnetosomes).
3 - 3 Membranes
Learning Objectives
After reading this section, students will...
- Know that membranes are highly conserved structures found in all living cells and be able to describe their structure.
- Understand the stabilizing forces of membranes.
- Be able to describe the functions of membranes.
- Be able to describe the function and location of the periplasm.
- Be able to describe the function of the cell wall and the two common cell wall structures.
General properties
Membranes define life. The cytoplasmic membrane immediately surrounds the inside of the cell and is perhaps the most conserved structure in living cells. Membranes are thin structures, measuring about 8 nm thick and every living thing on this planet has some type of membrane. They are the major barrier that separates the inside the cell from the outside and allow cells to selectively interact with their environment. Membranes are highly organized and asymmetric. This asymmetry comes from the fact that the membrane that faces the environment performs very different functions than does the side that faces the cytoplasm. Membranes are also dynamic, constantly adapting to changing environmental conditions.
Physical structure
Membranes are composed of lipids and proteins, with about half of all bacteria also containing hopanoids (see section 2-6). Proteins are more numerous in bacterial membranes than in eukaryotic membranes. Bacteria, in general, only contain a single membrane in contact with the cytoplasm, and this has to carry out all the functions of the cell. In eukaryotes, these functions are divided amongst the cytoplasmic membrane and the other organelles.
Some membrane proteins span the membrane (integral or intrinsic membrane proteins) while others sit on the outside or the inside (extrinsic or peripheral membrane proteins). Peripheral membrane proteins easily wash off the membrane using a salt solution, while intrinsic membrane proteins can only be released from it using detergents. Membrane proteins may move within the membrane plane, or the cell may anchor them to structures in or near the membrane.
Much of the general behavior of membranes is dictated by the behavior of lipids in water. Because phospholipids are amphipathic, they congregate when placed in an aqueous environment, similar to how oil will separate from water. This aggregation occurs in a particular way such that the hydrophilic portions face the water and the hydrophobic portions are buried inside. Under the cell's direction, lipids are organized into a bilayer, where there are two sheets of lipids oriented so that the hydrophobic faces of each sheet face each other, as shown in Figure 3.12. Lipid bilayers can be almost any size and can form vesicles spontaneously in water. In the cell, however, the synthesis of membranes is performed by specific enzymes and is tightly controlled.
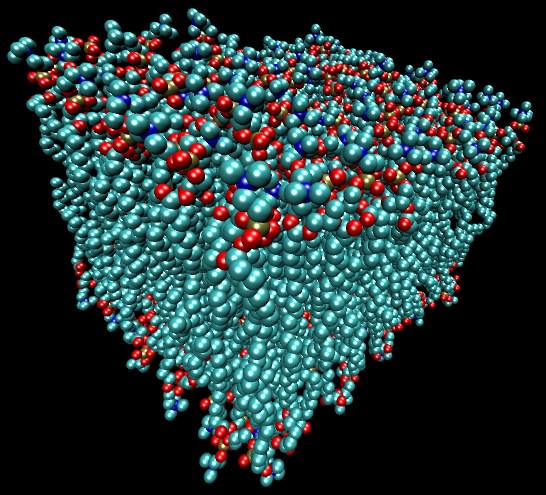

Figure 3.12. A model of a lipid bilayer. A space-filling representation of a lipid bilayer as developed by H. Heller et al. (J. Phys. Chem. 1993. 97:8343-60).
The cytoplasmic membrane is held together by several forces. Hydrophobic interactions between the alkyl chains of neighboring lipids are a major component of membrane stability. Hydrogen bonds between lipids and between membrane proteins and lipids also hold a membrane together. Further stability comes from negative charges on proteins that form ionic interactions with divalent cations such as Mg+2and Ca+2 and the hydrophilic head of lipids.
The membrane is fluid and has the consistency of a light-grade oil. It has been termed a fluid mosaic: "fluid" because the lipids are free to move about on each side of the membrane and "mosaic" because there is a definite pattern to it. Lipids do not generally switch from outside to inside or vice versa because it is energetically demanding to move the hydrophilic group through the hydrophobic core of the bilayer.
Membranes are a selective barrier
The concentration of solutes, sugars, and most ions is generally much higher within the cell than outside. A fundamental principle of nature is that different concentrations of a given solute tend to equilibrate across a boundary due to diffusion. The hydrophobic region of the membrane provides a critical function: it prevents this diffusion for polar molecules. Therefore, compounds such as amino acids, organic acids, and inorganic salts must be specifically transported across the membrane by proteins, and once inside, these molecules cannot escape. The cell can therefore control the nature and amount of these compounds that enter or leave the cell.
For molecules that are soluble in both the lipid membrane and the surrounding aqueous environment, the law of simple diffusion directs transport. The membrane is not a barrier for such molecules. These types of molecules are uncommon since solubility in both a hydrophobic and a hydrophilic environment is unusual. There is no transport protein for such compounds, so there is no specificity of control or energy cost. The cell cannot create a concentration gradient of these molecules.
There are three basic types of transport systems:
- Facilitated Diffusion
- Group Translocation
- Active Transport
Many of the proteins in the membrane function to help carry out selective transport, particularly of polar compounds. These proteins typically span the entire membrane, making contact with the outside environment and the cytoplasm. They often require the expenditure of energy to help compounds move across the membrane, though cells can also use concentration gradients of these compounds to generate energy, as described below
Facilitated diffusion
This process involves a protein that binds the molecule to be transported and physically moves that compound through the membrane. The molecule's binding to the protein causes a conformational change causing the molecule to face the opposite side from where it was. As shown in Figure 3.14, facilitated diffusion is therefore specific because a protein must bind the molecule. However, these small molecules are readily moved in and out of the cell, so a gradient cannot be formed, nor is energy required.
One example of a class of proteins that carry out facilitated diffusion is aquaporins. Water can diffuse relatively slowly across the membrane, but it was a matter of debate whether this was fast enough for cellular processes. Studies of adaptation to osmotic stress suggested that passive diffusion of water was too slow to explain the rapid changes observed in some bacteria. A search began for the elusive protein that was carrying out this process. Peter Agre discovered the first water channel in 1988. While first described in eukaryotic cells, many living systems are now known to contain them, and the term aquaporins was coined to describe them. Over 100 aquaporins are present in bacterial systems, and while these do not share a high degree of sequence homology with eukaryotic aquaporins, they share a significant structural similarity. There are two large classes of aquaporins: those that only allow water to flow through them and those that will also transport glycerol and a few other small uncharged molecules. Aquaporins are not present in every microorganism; some species appear to lack them. In these cases, the microbe may rely solely on diffusion across the membrane, or its aquaporin has a novel sequence.
Aquaporins (Figure 3.13) are intrinsic membrane proteins of about 23 kDa that contain six α-helices arranged around a central core channel. Each aquaporin will associate with three others to form a homotetramer in the membrane. The central channel is narrow and contains a conserved structure that can selectively allow only the passage of the desired molecule. In this way, water can rapidly flow into and out of the cell, speeding up its transport through the membrane.
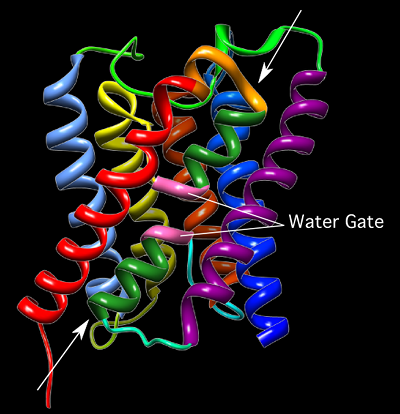
Figure 3.13. Aquaporin. A ribbon diagram of the 23 kDA Aquaporin from E. coli. Each of the 6 α helices that line the water channel are shown. The arrows indicate the path of water through the protein. The two sequences of 3 amino acids (Asparagine-Proline-Alanine) are shown in pink. This area is thought to be important in discriminating, only allowing water molecules to pass through the pore. Thus, serving as a gate.
While aquaporins are essential to cell function, they also create a serious problem. The inside of the cell is full of many types of solutes: proteins, nucleic acids, other small molecules, and ions. In comparison, the outside environment, in most cases, is very dilute. Because of this, there is a higher concentration of water outside the cell than inside the cell. Nature hates imbalances such as this, and to correct the issue, water tends to flow into the cell by a process called osmosis. Osmosis causes high pressure against the cell membrane. This pressure would rapidly cause lysis of most cells, and one of the major purposes of the peptidoglycan of the cell wall (discussed in a coming section) is to prevent the cell membrane from bursting. Figure 3.14 shows an animation of facilitated diffusion.
Figure 3.14. Facilitated Diffusion. An animation of the migration of solutes in and out of the cell as facilitated by a protein. Notice that this mechanism does not lead to a solute concentration inside the cell that is higher than outside. Rather, it leads to an equilibrium of that solute across the gradient.
Group translocation
In this process, a protein specifically binds the target molecule and transports it inside the cell while simultaneously modifying it chemically. Most group translocations require energy and tend to be unidirectional, unlike facilitated diffusion. The substrates of catabolic pathways, such as sugars, are sometimes transported by group translocation. Group translocation is an efficient way to both bring substrate into the cell and begin the breakdown process. Figure 3.15 shows an animation of group translocation.
Figure 3.15. Group Translocation. An animation of group translocation. The glucose molecule that is being transported into the cell is modified by the addition of a phosphate from phosphoenolpyruvate to form glucose-6-phosphate.
Active transport
In active transport, energy is expended to transport small molecules without chemically altering the substrate. The process is efficient enough to cause the internal concentration in the cell to reach many times its external concentration. Active transport proteins are molecular pumps that expend energy to pump their substrates against a concentration gradient. This energy comes in two forms: ATP and ion gradients. In ATP-based active transport, the transport proteins couples ATP hydrolysis to the movement of the small molecule across the membrane. The animation in Figure 3.16 shows an animation of an ABC transporter. One large group of proteins involved in this type of transport is the ATP binding cassette (ABC) transporters. ABC transporters are present in all living species, with 80 identified in the E. coli genome and 48 in the human genome. An example of an ABC transporter is the maltose-binding protein of E. coli.
Figure 3.16. The Maltose Transporter, an ATP Binding Cassette Transporter. Three proteins are involved in the maltose transporter. A maltose-binding protein in the periplasm and two membrane-bound transporter proteins. One serves at the gate, while the other binds and hydrolyzes ATP. The hydrolysis of ATP to ADP changes the conformation of the gate, allowing the passage of maltose into the cell.
Ion gradient active transport uses the energy of one chemical gradient, that of the specific ion, to drive the creation of a different gradient, the uptake of the small molecule. The ion gradient that supports the work is at a higher concentration on one side of the membrane than the other. The transport protein both binds its small molecule to transport and provides a gateway for this ion to fall down its concentration gradient. When the ion moves through its gateway, it causes a conformational change in the protein, and this change helps to transport the target small molecule into the cell.
Active transport proteins may be highly specific for only one molecule or may be able to carry a class of chemically related molecules. An example of a more general transport protein is the branch chain amino acid transporter of Pseudomonas aeruginosa, which transports leucine, valine, and isoleucine. Table 3.1 summarizes the various properties of transport mechanisms.
Table 3.1. Properties of various transport systems
| Property | Passive Diffusion | Facilitated Diffusion | Active Transport | Group Translocation |
| Carrier Mediated | - | + | + | + |
| Concentration Against Gradient | - | - | + | Not Applicable |
| Specificity | - | + | + | + |
| Energy Expended | - | - | + | + |
| Solute Modified During Transport | - | - | - | + |
A comparison of the methods for transporting molecules through the membrane and into the cell.
Membranes can help generate energy
Many cells use respiratory processes to obtain their energy. During respiration, organic or inorganic compounds that contain energy are oxidized, releasing electrons to do work. In many microorganisms, these electrons find their way to the membrane, passing through a series of electron carriers, as shown in Figure 3.17. During this operation, protons move outside the cell. This movement creates a gradient of protons across the cell membrane, energizing it in a fashion similar to charging a battery. The energy of this gradient can then be used to do work directly, a process known as the proton motive force, or can be channeled into a special protein known as ATP synthase. ATP synthase can convert: ADP to ATP, and the ATP can itself do work.
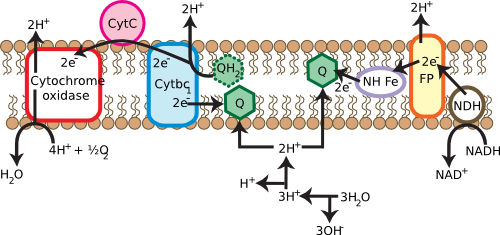
Figure 3.17. Generating the proton motive force. Membranes are critical in many cells for the generation of usable energy. The cartoon shows the various membrane proteins involved in converting high-energy electrons from photoreceptors into useful energy. They do so by forming a proton gradient across the cell membrane, termed a proton motive force, which is in turn used by other proteins to synthesize ATP. This will be discussed in greater detail in the chapter on metabolism.
The prokaryotic cells performing photosynthesis also have a membrane system that does a very similar process. Light excites electrons found in pigmented proteins in the membrane, and the electrons again move through a series of electron carriers. As above, a proton motive force is generated and used to synthesize ATP. The chapters on metabolism and photosynthesis discuss the specifics of these systems.
Membranes are also important in their own synthesis and can fold inward for specialized functions
Membranes also contain specialized enzymes that carry out certain biosynthetic functions. For example, the last few steps of lipid synthesis take place inside the membrane. Another example is cell wall synthesis and assembly. Much of the synthesis of cell wall monomers occurs there, and the stitching together of the cell wall polymer takes place while it is anchored to the membrane. In addition, any cellular protein that carries out its function outside the cell membrane (such as outer membrane and extracellular proteins) must pass through that membrane. During their synthesis, the cytoskeleton guides the ribosome to the cytoplasmic face of the membrane, and the growing peptide chain is synthesized directly into the lipid bilayer. Integral membrane proteins then fold up and stay in the membrane while extracellular proteins move through the membrane and take on their final shape on the other side.
Infoldings of the membrane are found in some photosynthetic bacteria. These bacteria use pigments in their membranes to capture light energy. Under low light, they need to increase the surface area to catch more light. They cannot make the membrane thicker, but they can increase the surface area by creating regions where the membrane folds into the cytoplasm. These invaginations are still attached to the cytoplasmic membrane and a picture of such structures, termed the intracytoplasmic membrane in the case of Rhodobacter sphaeroides, is shown in Figure 3.18.
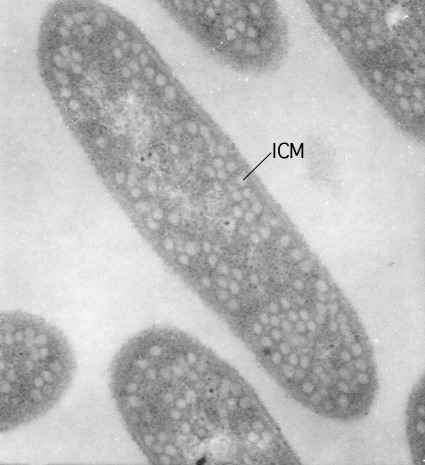
Figure 3.18. The intracytoplasmic membrane of Rhodobacter sphaeroides.. This electron micrograph shows the complicated infolding of the cytoplasmic membrane of R. sphaeroides when it is performing photosynthesis. This infolding creates a larger membrane surface area into which light-harvesting complexes can be inserted. Under low-light conditions many light-harvesting complexes are needed to capture the small number of photons striking the microbe. (Source: Samuel Kaplan, University of Texas - Houston Medical School)
Key Takeaways
- Membranes are some of the most conserved structures in biology. They are composed of mostly lipids, with some proteins and polysaccharides. Their characteristics are dictated by the amphipathic nature of lipids, containing a hydrophilic exterior and a hydrophobic core.
- Membranes serve as a selective barrier, preventing the passing of all but very non-polar molecules.
- To facilitate transport, membranes will contain transport proteins that move selected molecules across.
- The methods used by transport proteins can be classified as facilitated diffusion, group translocation and active transport.
- Group translocation and active transport concentrate molecules against a gradient and require energy.
- Membrane are also involved in the generation of energy and participate in their own synthesis.
3 - 4 The Bacterial Cytoskeleton
Learning Objectives
After reading this section, students will...
- Know that bacterial cells have cytoskeletal proteins.
- Know the role of MreB and crescentin in bacterial cell shape.
- Be able to describe how cytoskeletal proteins that control where FtsZ forms for cell division.
Scientists long thought that only eukaryotic cells had a cytoskeleton to perform the functions of movement of organelles, the beating of cilia, chromosome separation, and phagocytosis. Bacteria and Archaea were small enough that this kind of structure was undetectable and thought unnecessary. With the advent of whole-genome sequencing, researchers were surprised to find that microbes had homologs of all known eukaryotic cytoskeletal genes and proteins. Scientists are just beginning to understand the role of these proteins in the cell, with three of the best-understood proteins being, FtsZ, MreB, and crescentin. This section will describe how these proteins play a role in dictating cell shape, motility, chromosome separation, and cell division.
Cell Shape
MreB, as it is called in E. coli, is a homolog of actin, a eukaryotic protein involved in muscle contraction. MreB is a multi-functional protein, being important in forming cell walls (in rod-shaped cells), motility, chromosome segregation, cell-polarity (telling one end of the cell from another), and providing stability to the spore coat in Streptomyces species.
During its synthesis, the cell wall has to be synthesized in a distinct pattern. Even in the instance of bacteria that lack cells walls, cell shape is constant. In all these cases, the cytoskeleton plays a major role in helping to define the shape of the cell. Some of the most compelling evidence comes from mutants of MreB. MreB is a cytoskeletal protein found in most rod-shaped cells. MreB is an esential protein, as shown by experiments where inactivation of MreB (or similar genes in other bacteria) will cause the cells to lose their shape and become spherical, eventually becoming non-viable.
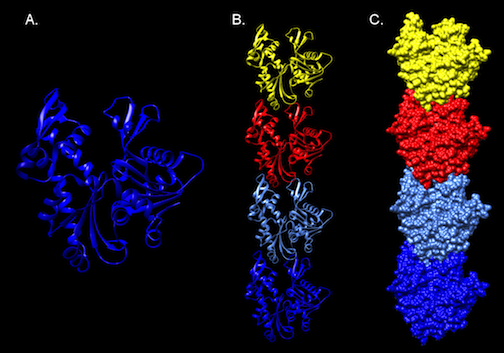
Figure 3.51. Molecular Structure of MreB. The crystal structure of MreB from Thermotoga Maritima. Panel A. A ribbon diagram of a monomer of MreB. A representation of part of a polymer of MreB as a ribbon diagram (Panel B), and as surface model (Panel C).Note the compact, building-block like structure of the protein, which can assemble into long filaments. Structure developed from the pdb file (1jce) as reported by Fusinita van den Ent, Linda A. Amos and Jan Löwe
MreB has a crystal structure that resembles actin (Figure 3-51), and it assembles into actin-like filaments. The latest research appears to show that MreB filaments assemble just inside the cell membrane. Initial observation of MreB in cells suggested the filament formed a helical structure that wound the length of the cell. Further inquiry has contradicted this picture with MreB forming strips instead of hoops inside the cell.
Crescentin (CreS) in Caulobacter crescentus forms a filament at the short axis of curved cell. This protein belongs to another type of cytoskeletal protein class called intermediate filament-like proteins (IF-like proteins). Scientists first described them in eukaryotes, but homologs are present in all domains of life. CreS is responsible for this curved shape, as CreS deficient mutants form straight rods. CtpS, a novel filament-forming enzyme, has also been discovered to be important in this process. CtpS and CreS interact, and regulation of this process dictates the curvature of the cell as it grows. Figure 3-52 shows various roles of cytoskeletal proteins in cell shape.
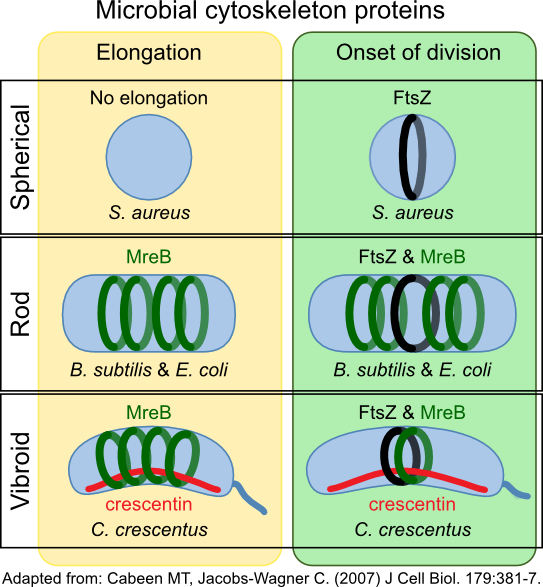
Figure 3.52. Cytoskeleton proteins and cell shape. The cytoskeleton proteins MreB and crescentin (CreS) help determine the shape of the cell. MreB forms hoops inside the cell membrane, directing cell wall synthesis. CreS accumulates on one side of the cell in Caulobacter species. This bends the cell, giving them their characteristic vibrio shape.
Other IF-like proteins have also been discovered in Helicobacter pylori. Four coiled-coil proteins (Ccrp48, Ccrp59, Ccrp1142, and Ccrp114) are essential for maintaining the spiral shape of the microbe. Deletion of any one of them will result in a fraction of the cells having a straight morphology. Scy (Streptomyces cytoskeletal element) is important in direction growth at the tips of hyphae of Streptomyces species and in determining new branch formation. A second IF-like protein in Streptomyces, FilP, is important in the stability of filaments in hyphae formation and for correct DNA segregation.
Motility
MreB has also been shown to be important in motility in some species. In Myxococcus xanthus, MreB and AglZ are important for forming a cytoskeleton onto which the gliding motility apparatus assembles. AglZ (another IF-like protein) assembles at the leading pole, migrates down the cell, and disassembles at the opposite pole. It seems to be forming a track to which adhesion proteins bind. These adhesion proteins then also interface with an environmental surface. The movement of the AglZ track down the cell thus causes the movement of the cell across the surface.
Chromosome separation
Three sets of proteins mediate separation of the chromosome during active growth by forming a partitioning system:
- A cytoskeletal proteins that hydrolyzes nucleotide triphosphates and provides energy, an NTPase
- A DNA-binding protein that complexes with the DNA
- A centromere like structure that is close to the origin of replication
Several classes of NTPases have been discovered that provide energy for chromosome separation (ParA, ParM, or TubZ). They are structurally similar but do not share significant sequence homology. In addition, the mechanisms investigated in various species appear to use different processes to power the migration of the DNA.
Centromere-like sites were first described in Bacillus subtilis (parS) and found to be bound by a DNA binding protein (ParB). Bioinformatic analysis of 400 bacterial species has shown the presence of parS sites in 69% of bacteria, making it the best conserved DNA sequence ever described in bacteria. These parS sites are located close to the origin of replication.
The arrangement of the parS site, the involvement of a DNA-binding protein, and an NTPase suggests the following mechanism: After replication, the DNA-binding protein attaches to the centromere site. Some time later, given an appropriate signal, the NTPase activates and drives the separation of the two chromosomes, using nucleotide triphosphates to power the process. The exact details of how this process proceeds are not clear.
Cell division
The cytoskeleton of bacteria also has a major role in the organization and implementation of cell division. An essential protein in this process is Fts. Fts stands for filamentous temperature-sensitive since mutants in this gene cause cells to be unable to divide and end up making long filaments in culture. Deleting this gene is lethal to the cell, and FtsZ is found in virtually all Bacteria and Archaea, with just a few exceptions.
After completion of chromosome replication and separation (directed by MreB), FtsZ proteins form a cytokinetic ring near the midplane of the cell. The point of formation of this ring is of great interest to the cells since forming it in the wrong place will cause an aberrant cell division. Thus, this process is tightly regulated by a whole series of proteins, Figure 5.53.
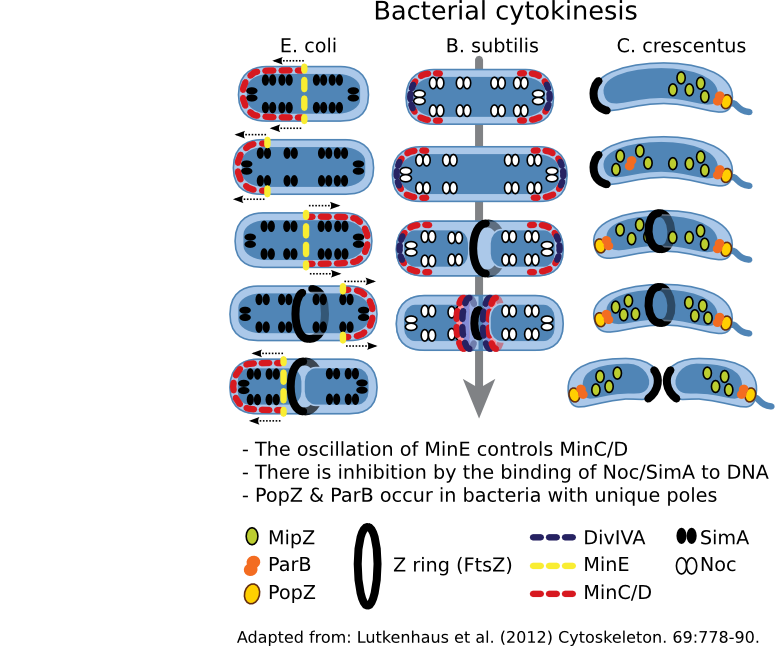
Figure 3.53. Regulation of cell division. The cytoskeleton in most bacteria directs cell division using FtsZ. This protein forms a Z ring that contracts, with concomitant hydrolysis of ATP, thus dividing the cytoplasm and directing cell wall synthesis. Where the Z ring forms is tightly controlled by the cell, but the mechanism used varies depending upon the bacterial species.
The Min system regulates the position of the Z ring in many bacteria so far examined. MinC is an antagonist of FtsZ and prevents Z ring formation. MinD, a membrane-bound protein, binds to MinC and localizes it to the membrane. In places where the concentration of MinC/D are high, FtsZ is unable to polymerize and form a Z ring. The presence of MinC/D is regulated by MinE, which binds to MinD and activates its ATPase activity, causing MinD to dimerize. In this conformation, MinD is knocked off the membrane, also removing MinC. A traveling wave of minE/D/C oscillates from end-to-end, resulting in the lowest concentration of MinC/D being at the center of the cell. Thus the Z ring forms in the center.
A second system is also found in many bacteria that employs nucleoid occlusion (NO). NO systems contain a protein (For example, SlmA in E. coli and Noc in B. subtilis) that binds to the DNA and, when bound to its site, will antagonize FtsZ polymerization. These sites are scattered throughout the chromosome but nearer the origin. Therefore, FtsZ ring formation cannot occur in the presence of DNA.
Both Min and NO systems suggest a model that contains a limited amount of FtsZ in the cell. However, Z-ring formation is promiscuous, and Min and NO prevent inappropriate ring formation at the cell poles or near the nucleoid. The absence of Min and the segregation of the chromosomes creates a low inhibition zone near the middle of the cell and allows Z-ring formation.
While Min and Slm/Noc have been found in a majority of bacteria, there are exceptions -- notably Caulobacter species. This genus lacks Min and Slm analogs and contains a unique system, where spatial the regulation system appears to be a hybrid of the Min and NO systems. A MinD-like protein, MlpZ, is recruited to the pole by ParB, bound near the origin of replication. Interaction with ParB dimerizes MlpZ, a form that antagonizes FtsZ. This dimerization is transient, and MlpZ eventually will decay into monomers. Thus a gradient of MlpZ is set up, whose concentration is dependent upon the nucleoid. When the chromosome replicates and segregates, a bipolar gradient is formed, with the low point being in the center of the cell where the Z-ring forms.
Key Takeaways
- Bacteria have cytoskeletal proteins.
- MreB and crescentin are important in determining the shape of the cell.
- FtsZ forms a contractable ring that divdies the cells. The location of this ring is regulated by nucleoid exclusion and the Min system in the microorganisms studied thus far.
3 - 5 The cell wall
Learning Objectives
After reading this section, students will...
- Be able to describe the function and location of the periplasm.
- Be able to describe the function of the cell wall and the two common cell wall structures.
The periplasm is between the cytoplasmic and outer membranes in Gram-negative bacteria
The periplasm is found in Gram-negative bacteria and is the space in between the cytoplasmic and outer membranes. (Some feel a periplasm-like compartment is also present in Gram-positive bacteria between the cytoplasmic membrane and the peptidoglycan.) The periplasm is filled with water and proteins and is therefore somewhat reminiscent of the cytoplasm. However, pools of small molecules in the periplasm are not like those in the cytoplasm because the membrane prevents the free exchange between these two compartments. Also, the proteins found in the periplasm are distinct from those in the cytoplasm. They are specifically guided to this site during translation through specific signal sequences typically near their N-termini. Table 3.2 lists some examples of these proteins.
The peptidoglycan shell that provides the strength to prokaryotic membranes is also found in the periplasmic space of Gram-negative bacteria. In contrast, in Gram-positive bacteria, peptidoglycan provides the outside border to the periplasm.
Table 3.2 Different types of periplasmic enzymes and their role in the cell
| Enzyme Type | Examples | Function |
| Hydrolytic enzymes | phophatases | Degrading phosphate-containing compounds. |
| proteases | Degrading proteins and peptides. | |
| endonucleases | Degrading nucleic acids. | |
| Binding proteins | sugars, amino acids, norganic ions, vitamins | Binding substrates and docking with transport protein in membrane. |
| Chemoreceptors | Chemotaxis, ermination | Sensing the environment and changing cell behavior in response. |
| Detoxifying enzymes | β-lactamase | Degrading penicillin and related compounds before they get into the cell. |
Periplasmic enzymes have several main functions, detecting nutrients in the environment, degradation of polymers, and protection from harmful compounds.
The cell wall surrounds and holds in the microbe
This section will restrict itself to the bacterial cell wall. We will compare this to archaeal cell walls at the end of the chapter. The cell wall is essential to the survival of most microorganisms. Many microbes live in relatively dilute environments, and the wall's most important function is to prevent the cell from bursting due to osmotic stress. The cell wall also determines the shape of the cell. Any cell that has lost its cell wall, either artificially or naturally, becomes roughly spherical and lyses due to osmotic pressure unless placed in certain concentrated solutions. Finally, the cell wall helps to support any structure that penetrates from the cell out into the environment.
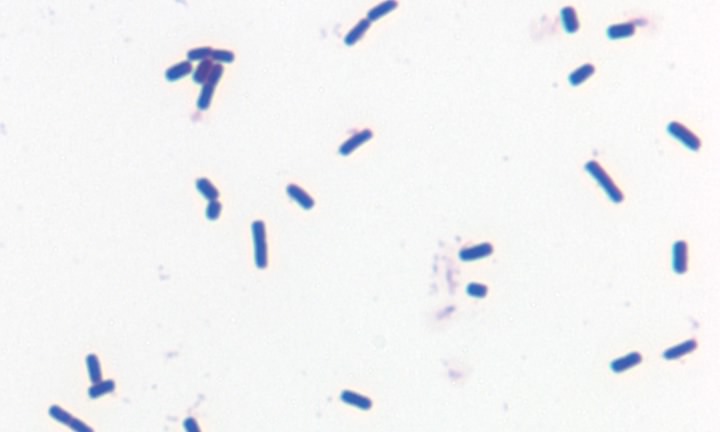
Figure 3.19. A gram-positive bacterium. Gram stain of the gram-positive bacterium Bacillus cereus
The structure and synthesis of prokaryotic cell walls is unique, and many compounds found in the bacterial cell wall are found nowhere else in nature. It is true that plants also make cell walls, but they are chemically and structurally different. Investigations have revealed two basic types of bacterial cell wall structures: Gram-positive and Gram-negative. These two classes of bacterial cells look very different following staining with the Gram stain, and this has been a standard test for identification of bacterial species. Figures 2-19 and 2-20 show Gram stains of Gram-positive and Gram-negative bacteria, respectively.
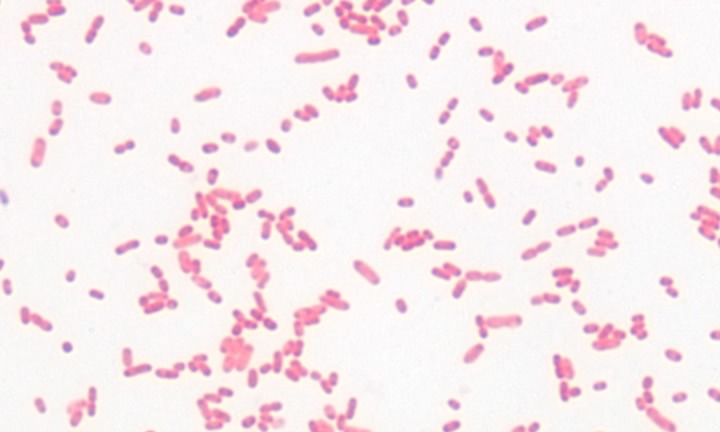
Figure 3.20. A gram-negative bacterium. A Gram stain of the gram-negative bacterium Serratia marcescens
When Hans Christian Gram developed the Gram stain in 1884, the molecular basis of the stain was unknown. Bacteriologists understood very little about bacteria in general. He just empirically determined that when bacterial smears were run through a four-step staining procedure using two different dyes, some cells retained the first dye and stained purple, while others only retained the second dye and stained pink. Years later, bacteriologists discovered that this differential reaction relates to the cell wall, as shown in Figure 3.21.
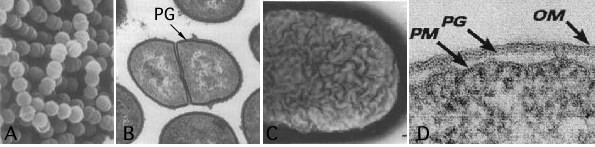
Figure 3.21. A comparison of the ultrastructure of gram-positive and gram-negative cells. The different Gram reactions occur because of structural differences between the bacterial cell walls. Gram-positive cells (Group B streptococci) appear smooth in a scanning electron micrograph (A) and are composed of a single layer of peptidoglycan (B). Gram-negative cells (E. coli) have an undulating surface and have three layers (C and D). (Sources: S. H. Pincus, et al.1992. J. Bacteriol 174:3739-3749 [panels A and B]; M. E. Bayer and C. C. Remsen. 1970. J. Bacteriol. 101:304-313 [panel C]; T. J. Beveridge. 1999. J. Bacteriol. 181:4725-4733 [panel D])
As shown in Figure 3.21, the Gram-negative cell has an additional layer, and the outside of the cell appears convoluted when compared to the Gram-positive cell. The Gram-positive wall is much thicker than is the Gram-negative wall, and its external appearance is smoother. Gram-positive and Gram-negative cells do share one thing in common that is unique to bacteria - peptidoglycan. We will talk about the structure of this and then examine the various structures found in each cell wall type.
Peptidoglycan is a thick rigid layer composed of an overlapping lattice of two sugars, N-acetyl glucosamine (NAG) and N-acetyl muramic acid (NAM), that are cross-linked by amino acid bridges, as shown in Figure 3.22. The exact molecular makeup of these cross-bridges is species-specific. NAM is only found in the cell walls of bacteria and nowhere else. Attached to NAM is a side chain generally composed of four amino acids. In the best-studied bacterial cell walls (E. coli), the cross-bridge is most commonly composed of L-alanine, D-alanine, D-glutamic acid, and diaminopimelic acid (DPA).
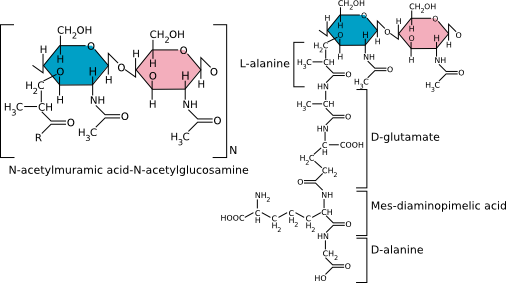
Figure 3.22. The chemical structure of peptidoglycan. The generalized peptidoglycan monomer showing the two sugars that make up the backbone. The R group consists of four amino acids, with the best-studied cell walls containing L-alanine, D-alanine, D-glutamic acid and diaminopimelic acid.
Note that peptidoglycan contains D-amino acids, which are different than the L-amino acids found in proteins. D-amino acids have the identical composition as L-amino acids but are their mirror images. The use of D-amino acids is unusual in biology, and bacteria have enzymes called racemases to convert between D and L forms specifically for this use.
The NAM, NAG, and amino acid side chain form a single peptidoglycan unit that can link with other units via covalent bonds to form a repeating polymer. Covalent bonds between cross-bridges further strengthen the polymer, and the degree of cross-linking determines the degree of rigidity. Figure 3.23 shows an artist's rendering of what the structure might look like. In E. coli, the penultimate D-alanine of one unit links to DPA of the next cross-bridge. In some Gram-positive microbes, there is a peptide composed of various amino acids that link the cross-bridges. For example, in Staphylococcus aureus strains, five glycines make up the linker between peptidoglycan monomers. The sequence of these linkers varies considerably between species. The completed peptidoglycan layer forms a strong mesh analogous to a chain-link fence. The complete cell wall contains one or more layers of peptidoglycan, one atop the other, providing much of the strength of the cell wall.
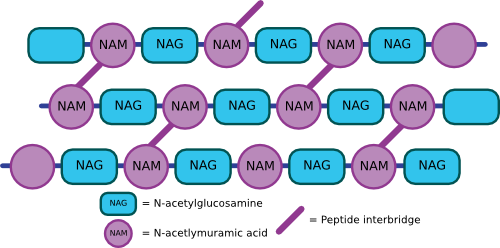
Figure 3.23. A cartoon of the peptiodglycan mesh. The peptidoglycan polymers then crosslink with other peptidoglycan chains to form a complex mesh that wraps the cell in a structure a kin to chicken wire.
While both Gram-negative and Gram-positive bacteria have peptidoglycan, their physical arrangement in the cell wall differs. In Gram-positive cells, the peptidoglycan is a heavily cross-linked woven structure that encircles the cell in many layers. It is very thick, with peptidoglycan accounting for 50% of the cell weight and 90% of the weight of the cell wall. Electron micrographs show the peptidoglycan to be 20-80 nm thick. In Gram-negative bacteria, the peptidoglycan is much thinner, with only 15-20% of the cell wall being peptidoglycan. In both cases, peptidoglycan is not a barrier to solutes, as the openings in the mesh are large enough for most molecules, including proteins, to pass through. Figure 3.24 shows a depiction of the Gram-positive cell wall.
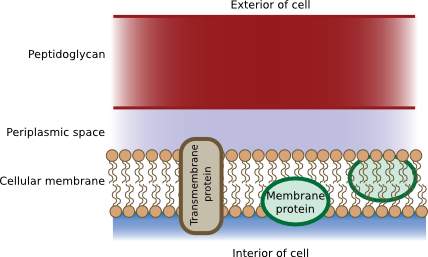
Figure 3.24. The Gram-positive cell wall. The cell wall is made mostly of peptidoglycan, interspersed with teichoic acid which knits the different layers together. The amount of crosslinking is higher and the wall is thicker than in gram-negative cell walls.
The Gram-positive cell wall
Gram-positive cells consist almost entirely of peptidoglycan, but teichoic acid is an important structure found in Gram-positive cell walls. It is a phosphodiester polymer of glycerol or ribitol joined by phosphate groups. Amino acids such as D-alanine are attached. Teichoic acid is covalently linked to muramic acid and stitches various layers of the peptidoglycan mesh together. Teichoic acid stabilizes the cell wall and makes it stronger. Figures 3.25 shows the chemical formula of teichoic acid.
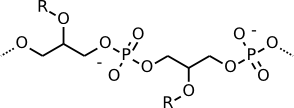
Figure 3.25. Teichoic acid. Teichoic acid is a long, thin molecule that weaves through the peptidoglycan.
Gram-negative cell structure
Gram-negative cell walls have a more complicated structure than that of Gram-positive organisms. Outside the cytoplasmic membrane is the periplasm, which contains the thin layer of peptidoglycan. The peptidoglycan in Gram-negative cells contains less cross-linking than in Gram-positive cells with no peptide linker. Covalently bound to the peptidoglycan is Braun's lipoprotein, which has a hydrophobic anchor that helps to bind the peptidoglycan to the outer membrane. Figure 3.26 shows the arrangement of the Gram-negative cell wall.
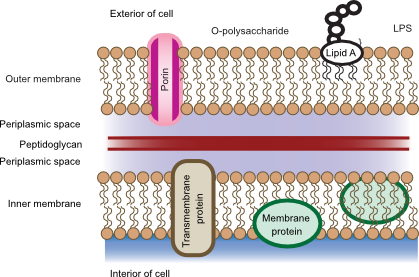
Figure 3.26. The Gram-negative cell wall. The cell wall in Gram-negative bacteria contains much less peptidoglycan and is surrounded by an outer membrane. There is much less crosslinking between the peptidoglycan. LPS is also present in the outer membrane and penetrates into the surrounding environment.
The outer membrane
The outer membrane of Gram-negative bacteria is another lipid bilayer similar to the cytoplasmic membrane, and contains lipids, proteins, and also lipopolysaccharides(LPS). It is a barrier to proteins and prevents enzymes secreted into the periplasm from floating away. The membrane has distinctive sides, with the side that faces the outside containing all the LPS. LPS is composed of two parts: Lipid A and the polysaccharide chain that reaches out into the environment. Lipid A is a derivative of two NAG units with up to 7 hydrophobic fatty acids connected to it that anchor the LPS in the membrane, as shown in Figure 3.27. Attached to Lipid A is a conserved core polysaccharide that contains KDO, heptose, glucose, and glucosamine sugars. The rest of the polysaccharide consists of repeating sugar units, and this is called the O-antigen. The O-antigen varies among bacterial species and even among various isolates of the same species. Many bacterial pathogens vary the makeup of the O-antigen to avoid recognition by the host's immune system.
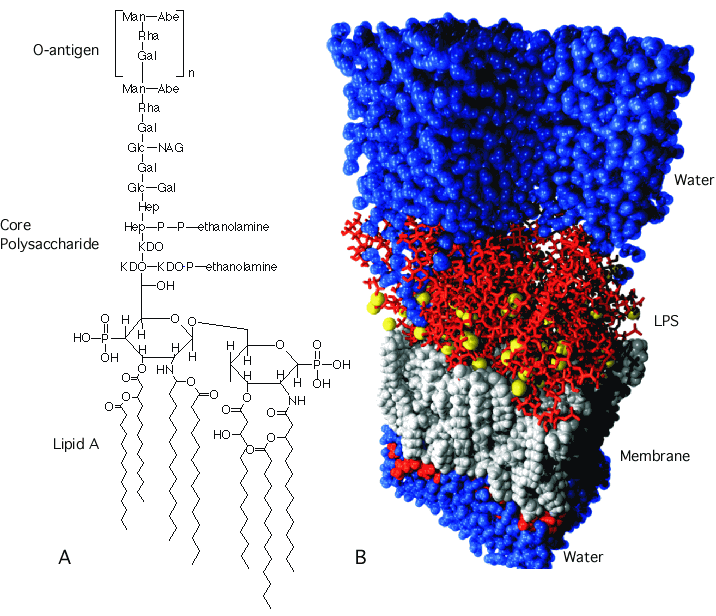
Figure 3.27. The structure of LPS. LPS is composed of three sections: the lipid A region, a conserved core polysaccharide, and a highly variable O-polysaccharide. (A) The chemical structure of LPS. (B) A molecular model of the outer membrane from Pseudomonas aeruginosa. Source (T. P. Straatsma, Pacific Northwest National Laboratory)
LPS confers a negative charge and also repels hydrophobic compounds, including certain drugs and disinfectants that would otherwise kill the cell. Some Gram-negative species live in mammals' gut, and LPS repels fat-solubilizing molecules such as bile that the gall bladder secretes. This repulsion enables these bacteria to survive in this environment. The O-antigen and other molecules on the outer membrane are targeted by certain viruses that infect bacteria as a means to identify the correct hosts for infection.
LPS is medically important because when LPS is released from bacterial cells it is toxic to mammals and is therefore called endotoxin. It creates a wide spectrum of physiological reactions including the induction of a fever (endotoxins are said to be pyrogenic), changes in white blood cell counts, leakage from blood vessels, tumor necrosis and lowered blood pressure leading to vascular collapse and eventually shock. At high enough concentrations the LPS endotoxin is lethal.
There are fewer total proteins and fewer unique types of proteins in the outer membrane than in the cytoplasmic membrane. Porins are essential components because of their role in the permeability of the outer membrane to small molecules. Porins form pores in the outer membrane large enough to allow the passage of most small hydrophilic molecules. Figure 3.28 shows the structure of a porin at the molecular scale. All known porins have a similar structure, with the protein containing a central channel that allows the passage of molecules. This common structure allows the migration of these molecules into the periplasmic space for possible transport across the cytoplasmic membrane. Some porins in the outer membrane are general, doing simple discrimination on size and charge, but having little substrate specificity. Examples include OmpF, which is selective for positively charged molecules, and PhoE, which is permeable to negatively charged molecules. Other porins are more specific. The best-studied is LamB, which recognizes the sugar polymer maltooligosaccharide and transports it through the outer membrane. Very large or hydrophobic molecules cannot penetrate the outer membrane, so the outer membrane serves as a permeability barrier to at least some molecules.
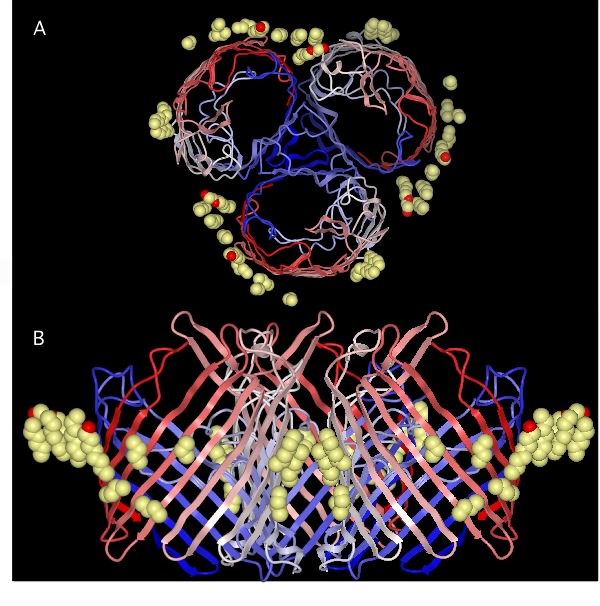
Figure 3.28. The molecular structure of a porin. The view in (A) is from the outside of the cell looking at the membrane surface. The view in (B) is the perspective from the side (i.e. from the membrane). The porin has three protein subunits and the actual pore is the central triangular area in the top panel formed by the three subunits.
Other types of outer membrane proteins are involved in various functions. OmpA in E. coli seems to connect the outer membrane to the peptidoglycan. Some pathogens contain outer membrane proteins that help them neutralize host defenses. Finally, all Gram-negative bacteria contain high molecular weight proteins involved in the uptake of large substrates such as iron-complexes and vitamin B12.
The differences between Gram-positive and Gram-negative bacterial cell walls greatly influence the success of the microbes in their environments. The thick cell wall of Gram-positive cells allows them to do better in dry conditions because it reduces water loss. The outer membrane and its LPS help Gram-negative cells excel in the intestines and other host environments. Table 3.3 summarizes the difference between Gram-negative and Gram-positive cell walls.
Table 3.3 Properties of cell walls
| Property | Gram-positive | Gram-negative |
| Thickness of wall | 20-80 nm | 10 nm |
| Number of layers in wall | 1 | 2 |
| Peptidoglycan content | >50% | 10-20% |
| Teichoic acid in wall | + | - |
| Lipid and lipoprotein content | 0-3% | 58% |
| Protein content | 0% | 9% |
| Lipopolysaccharide | 0 | 13% |
| Sensitive to penicillin | Yes | Less sensitive |
| Digested by lysozyme | Yes | Weakly |
A summary of the differences between Gram-positive and Gram-negative cell walls.
Some bacteria lack cell walls
For most bacterial cells, the cell wall is critical to cell survival, yet some bacteria do not have cell walls. Mycoplasma species are widespread examples, and some can be intracellular pathogens that grow inside their hosts. Cell walls are unnecessary here because the cells only live in the controlled osmotic environment of other cells. They could likely form a cell wall at some point in the past, but as their lifestyle became one of existence inside other cells, they lost the ability to form walls. Consistent with this very limited lifestyle within other cells, these microbes also have very small genomes. They do not need the genes for all sorts of biosynthetic enzymes, as they can steal the final components of these pathways from the host. Similarly, since their intracellular environment is completely predictable, they can also do without genes encoding many different pathways for various carbon, nitrogen, and energy sources. Because of the absence of cell walls, Mycoplasma have a spherical shape and are quickly killed if placed in an environment with very high or very low salt concentrations. However, Mycoplasma do have unusually tough membranes that are more resistant to rupture than other bacteria since this cellular membrane has to contend with the host cell factors. Sterols in the membrane contribute to membrane durability by helping to increase the forces that hold the membrane together.
Key Takeaways
- Outside the cell membrane is the periplasm, where enzymes often accumulate that perform degradative and transport functions.
- Outside the periplasm in most microbes is the cell wall. The cell wall is important in giving the cell its shape and preventing the cell membrane from rupturing due to the high osmotic pressure it is under.
- Two general types of cell wall structure are found in bacteria, Gram positive and Gram negative.
- Gram-positive cells have a thick peptidoglycan layer surrounding the cell membrane.
- Gram-negative cells have a thin peptidoglycan layer that is surrounded by a second membrane structure, the outer membrane.
- Finally, there are bacteria that have no cell walls.
3 - 6 The cell surface extends into the environment
Learning Objectives
After reading this section, students will...
- Know the structure and functions of flagella, pili and fimbriae.
- Be able to explain the composition and purpose of capsules and slime layers.
Surface structures are typically attached to a membrane, and extend into the environment. Important structures include flagella, pili, fimbriae, and glycocalyx. These protrusions and surfaces interact with the environment around the microorganism and therefore are pivotal in how the microbe sees the world and how we see the microbe.
Flagella are one type of structure used for motility
In many bacteria, flagella are responsible for motility in liquid. There is also a loose correlation between cell shape and the presence of flagella. Almost all spirilla, half of all rod-shaped bacteria, and only a few of the cocci are motile by flagella. One rationale for this correlation might be that spherical cells such as the cocci simply do not have the best geometry for directional movement by flagella, unlike more linear bacteria.
Flagella can be thought of as little semi-rigid propellers free at one end and attached to a cell at the other. Flagella are thin (20 µm) and long, with some having a length 2-3 times (about 10 µm) the length of the cell. Due to their small diameter, flagella cannot be seen in the light microscope unless a special stain is applied. Bacteria can have one or more flagella arranged in clumps or spread over the cell surface. Figure 3.29 demonstrates some of the more common arrangements.

Figure 3.29. Flagellar arrangements. A cartoon of several common flagellar arrangements.
Flagella are mostly composed of the protein flagellin, which is bound in long chains and wraps around itself in a left-handed helix as shown in Figure 3.30.
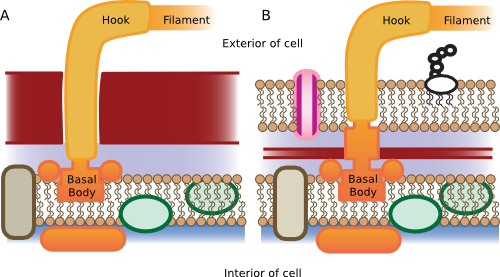
Figure 3.30. Flagella structure. Flagella are attached by a hook and rings that anchor it to the cell wall of the microorganism. In gram-positive bacteria (A) the rings are located in the cytoplasmic membrane and the flagella passes through the peptidoglycan to the outside environment. In gram-negative bacteria (B) there are additional protein rings in the outer membrane.
Flagella are organs of motility
The flagellum attaches to the cell through complex protein structures termed the hook and the basal body, which span the cell membrane. One ring in the basal body rotates relative to the other, causing the flagellum to rotate. The proton gradient, created by metabolism, provides the energy to drive this basal body (we will discuss this in much more detail in the metabolism chapter). In some fashion, the translocation of protons from outside to inside the membrane causes the rotation of the flagellum. In a sense, protons move through the wheel-like basal body (similar to a water wheel, except using protons), which causes the rotation of the assembly, including the flagellum. When E. coli is swimming through a solution, the flagella turn counter-clockwise and push the microbe. This behavior is termed smooth swimming. It is possible for E. coli to also reverse the direction of flagellar rotation, and when the flagella turn clockwise, they pull against the bacterial cell. Since E. coliis flagellated peritrichously (that is, at many positions), it is pulled in all directions and tumbles.
How fast do bacterial cells move? The average is 50 µm/sec, which is about 0.00015 kilometers/hr. speed may seem slow but remember their tiny size. Table 2.4 shows a better comparison and indicates that, relatively speaking, bacteria are faster than humans. Also, remember that this motility happens in water, which is much more viscous than air.
Table 2.4 Relative speeds of organisms
| Organism | Kilometers per hour | Body lengths per second |
| Cheetah | 111 | 25 |
| Human | 37.5 | 5.4 |
| Bacteria | 0.00015 | 10 |
Bacteria seem slow, until you consider relative size. Then they are quite fast.
If a flagellum breaks off, it is resynthesized until it reaches the appropriate length. This growth occurs from the tip. The flagellar filament is hollow, and flagellin monomers are passed up through this space until they reach the growing tip and are addadded to the structure.
Taxis and flagella
Typically microbes in aqueous environments continually move around looking for nutrients. Even microorganisms in the soil have uses and opportunities for movement. Sometimes this movement is random, but in other cases, it is directed toward or away from something. As a rough guide, bacteria want to move towards food or energy sources and away from toxic compounds. In other words, bacteria are capable of showing simple behavior that depends upon various stimuli.
There are several classifications of tactic responses, and the category is based upon the stimulus that the movement is responding to.
- chemotaxis - towards or away from a chemical stimulus
- phototaxis - towards light
- aerotaxis - towards or away from oxygen
The purpose of chemotaxis should be fairly evident: attraction to nutrients or avoidance of damaging compounds. A cell distinguishes chemicals that fall into either of these classes with receptor molecules near their surfaces that can tell if specific chemicals are in the environment. Many of these receptors are also involved in transport of their target molecules. Now it is trickier than it sounds. A cell does not merely want to sense the presence of nutrients but wants to move toward the highest concentration. It does this by an amazing process that is sort of a primitive memory, where it essentially asks if its receptors are binding more or less of the compound than they were a moment ago (a moment ago being about 200 milliseconds.). If they are binding more, the cell keeps swimming, but if they are binding less, it tumbles and then swims in a random direction. This tumbling is the direct result of E. coli's ability to reverse the rotation of its flagella.
Phototaxis is somewhat different, but again the cells move toward optimal levels of specific wavelengths of light. Rhodobacter sphaeroides (a photosynthetic microbe) performs phototaxis by a mechanism analogous to chemotaxis in E. coli, but there are important differences. R. sphaeroides contains only one polar flagellum, which rotates during a run toward light. If, during a run, light conditions worsen, the flagella stop rotating (instead of reversing its rotation). The microbe depends upon Brownian movement in the environment to turn it in a new direction.
Aerotaxis refers to the ability of some bacteria to respond to the presence of oxygen. It is mechanistically generally similar to what we previously described and depends on dissolved oxygen levels in the environment. Whether a bacterium swims toward it or away depends on the type of metabolism that it has.
Some bacteria move by gliding motility
Flagella are not the only means bacteria have developed for moving about the environment. A large collection of phylogenetically diverse bacteria have developed gliding motility, an energy-requiring process by which bacteria move smoothly over a solid surface. Gliding microbes tend to be predominately Gram-negative, but there are examples of Gram-positive bacteria. This type of motility is not as well understood as flagellar propulsion, and there appear to be several different mechanisms employed to accomplish it.
In one mechanism, the microbes use pili (type iv pili specifically) that are extended away from the cell and stick to the surrounding surface. The microbe then pulls itself toward the tethered end by retracting the pilus back inside the cell. By repeating this process, the cell drags itself along a surface. This type of motility is observed in a number of microorganisms including Pseudomonas aeruginosa, Neisseria gonorrhoeae, and Myxococcus Xanthus. Movement in this manner requires energy in the form of ATP.
There are also gliding bacteria that use other mechanisms. Some filamentous cyanobacteria seem to extrude an extracellular polysaccharide through small pores on the surface of the organism. Figure 3.31 depicts this type of gliding motility. Researchers think that the polysaccharide exiting from the cell propels the microbe along the surface by some means.
Figure 3.31. Gliding motility in Myxococcus xanthus.. Vegetative cells of a Myxococcus xanthus moving across an agar surface using gliding motility.
A third groups of microbes (Cytophaga and Flavobacterium) seem to use yet another mechanism that is dependent upon proton motive force. Observation of these microbes in the presence of tiny latex beads shows the beads moving along the microbe's surface directionally. The bead moves from one pole of the rod-shaped bacteria to the other, sometimes reversing direction. One model to explain this behavior is the concerted movement of cytoplasmic membrane proteins that are coupled to outer membrane proteins. These membrane proteins might form a kind of conveyor belt. The outer membrane proteins in this mechanism contact the environmental surface, and their movement propels the microbe forward.
Interestingly, Myxococcus xanthus uses several of these mechanisms in its motility. M. xanthus is a social predator. It glides around in large groups of cells secreting toxins and degradative enzymes that kill other microbes. The leftovers of these dead microbes then provide nutrients for the marauding Myxococcus. Investigations by Dale Kaiser and others have revealed that this microbe has two types of gliding motility, social and adventurous. Social motility occurs in groups, while adventurous motility involves single cells. It turns out that social motility is dependent upon pili, as described above, while adventurous motility involves the extrusion of slime from pores in the microbe's cell wall.
Pili
Pili and fimbriae are structurally similar to flagella and are composed of one or a few proteins arranged in a helical fashion. Figure 3.32 shows pili isolated from Neisseria gonorrhea. Each protein subunit assembles on the growing structure at the tip, as is the case with flagella. There are many genes necessary for the successful construction of pili. Their products might perform functions such as moving the structural proteins across the membrane, methylating the structural proteins, or retracting the pilus. The same is generally true for fimbriae.
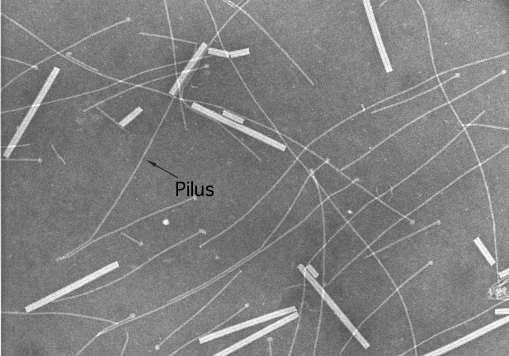
Figure 3.32. Pili and fimbriae. The pili shown in this micrograph are those of Neisseria gonorrhea with Tobacco Mosaic virus (the thicker structures) added as a size reference. The width of the Tobacco Mosaic virus is 0.018 µm. (Source: Katrina Forest, University of Wisconsin-Madison)
Fimbriae are found on many bacteria, are shorter and straighter than flagella, and more numerous. Not all bacteria synthesize them. Fimbriae do not function in motility but seem to be important in attachment to surfaces. Some microbes attach to hosts by fimbriae, and successful colonization of many surfaces depends on the ability to make fimbriae. Swarming microbes such as Myxococcususe them to sense the presence of similar microbes, which helps keep their "hunting packs" together.
Pili are longer than fimbriae, and there are only a few per cell. There are two basic functions for pili: gene transfer and attachment to surfaces. In gene transfer, the pilus captures a recipient bacterium, and DNA will be transferred from the donor to the recipient microbe. Conjugation, as this transfer process is known, is one explanation for the rapid spread of drug resistance in many different species of bacteria and is covered in depth in the Chapter on Genetics and Genomics.
Pili are also important for the attachment of many types of microorganisms to surfaces. For example, Neisseria gonorrhoeae, the causative agent of gonorrhea, has a special pilus that helps it adhere to the urogenital tract of its host. The microbe is much more virulent when able to synthesize pili.
Bacterial cells are often covered in glycocalyx
The general term for any network of polysaccharides or protein-containing material extending outside the cell is the glycocalyx. Many bacteria produce such a coating on the outside of their cell, and they come in two types: capsules and slime layers. The difference between the two is somewhat arbitrary. A capsule is closely associated with cells and does not wash off easily, while a slime layer is more diffuse and easily washes away. Figure 3.33 shows the capsule surrounding Klebsiella planticola.
Figure 3.33. Capsule surrounding cells of Klebsiella planticola.. The capsule (made of polysaccharide) in the figure is colorless and about the diameter of the cell. The background is darker.
There are many different types of proteins, polysaccharides, polyalcohols, and amino sugars in glycocalyx, and the exact makeup is species-specific. The structure can be thick or thin, rigid or flexible. Both Bacteria and Archaea are known to make capsules that will surround the cell.
There are several functions attributed to glycocalyx, one of which is to help cells attach to their target's environment. Streptococcus mutans produces a slime layer in the presence of sucrose. This forms a surface that allows other bacteria to aggregate on tooth surfaces and results in dental plaque. Vibrio cholerae, the cause of cholera, also produces a glycocalyx that helps it attach to the intestinal villi of the host. Glycocalyx can play other roles in pathogenesis as well. Bacteria that enter the body are always in danger of being attacked by phagocytes (host cells that protect you from invaders). Often the capsule makes it more difficult for phagocytes to attach to and engulf pathogens. As one example, Streptococcus pneumoniae, when encapsulated by a glycocalyx, can kill 90% of infected animals, while non-encapsulated forms cannot. In addition, capsules and slime layers are largely hydrophilic, so they can bind extra water in the environment and contribute to the protection of the cell from desiccation. Capsules and slime layers can also provide protection from the loss of nutrients by holding them within the layer. These extra layers coating the surface of the cell may also potentially mask viral receptors making it more difficult for viruses to attach. Many of these functions manifest themselves in biofilms. The glycocalyx is the glue that holds together whole communities of microorganisms. We will discuss biofilms in more detail in the chapter on Environmental Microbiology.
Key Takeaways
- Outside of the cell wall, bacteria contain structures that help them interact with their environment.
- For motility, microbes often use flagella, which can help them to move toward helpful environments and away from damaging environments.
- Other microbes can move using gliding motility, a method that depends upon pili or the excretion of slime.
- Pili can also be used for attachment to surfaces or as a method of exchanging genetic information.
- Fimbriae are shorter than pili, but can also serve at organs of attachment.
- Bacteria will also excrete a glycocalyx, a layer composed of protein or polysaccharide that helps the microbe stick to surfaces, and protect itself. Glycocalyx is important in the formation of biofilms.
3 - 7 Bacteria can exist in different cell states
Learning Objectives
After reading this section, students will...
- Be able to explain the structure and function of resting cells.
- Be able to explain why endospores are resistant to heat, chemicals and radiation.
- Be aware of different cells states into which microbes can differentiate.
For obvious reasons, we have focused on growing cells, but there are non-growing states of microbes that are important to both microbes and humans. In these states, termed spores and cysts, the cells remain dormant for long periods of time. Part of the relevance of these states is that the same properties that allow the cell to survive extended periods also happen to make the cells resistant to our typical efforts to kill them. Consequently, spores and cysts can thwart attempts to sterilize a sample. In this section, we examine some of the properties of these structures.
Spores and cysts
Spores and cysts are resting structures. That is, these states have very low to nonexistent rates of metabolism. They are common in organisms that live in soil and may need to survive some rough conditions such as lack of nutrients, high heat, radiation, or drying.
Sporulation is a unique developmental cycle. After a cell decides to sporulate, the creation of a different type of cell needs to take place, which requires turning on a large collection of genes in a tightly coordinated fashion. In addition, all of this expression must be complete before the microbe runs out of energy. There are several types of spores. Some are highly resistant structures that form under conditions of cell stress inside a supportive cell. These are called endospores. Others are part of the normal reproductive cycle, being created by the differentiation of a vegetative cell, and we will refer to these as spores. In this section, we talk generally about the structure of spores, and in the chapter on regulation, we will examine the regulation of sporulation.
Endospores
Endospores are refractile - light cannot penetrate them - so that they are easy to see in the phase microscope, and this makes it easy to detect them. Most endospores have no measurable metabolism and are a form of suspended animation. Endospores can survive for a very long time and then return to a growing state, a process termed germination. Endospores that were dormant for thousands of years in the great tombs of the Egyptian Pharaohs were able to germinate and grow when placed in appropriate medium. Several scientists have recovered viable endospores from bees trapped in amber that is 25-40 million years old. The microbe isolated was found to be most closely related to Bacillus sphaericus. There are even claims of endospores over 250 million years old being able to germinate when placed in an appropriate medium, but these claims still need to be verified. Endospores are everywhere, are easily dispersed throughout the environment, and can be difficult to remove. The anthrax scare of 2001 in the United States is ample evidence of the insidiousness of endospores and their impressive resistance.
The formation of an endospore is clearly a great advantage for these bacteria and enables them to endure extreme stress. At a later time, even much later, when conditions are favorable, they can reemerge and flourish. Endospores enable a species to spread easily from one suitable environment to another, and many endospore-forming bacteria are ubiquitous in the environment. Endospores are a particular problem in the food industry. Companies must take great care to ensure either the destruction of endospores or suitable preservation methods so that endospore-forming bacteria (and other microbes) cannot grow.
Endospores are resistant to heat (>100 °C), radiation, many chemicals (i.e. acids, bases, alcohol, chloroform), and desiccation. The mechanisms that account for this resistance include the impermeability of the endospore coat, the dehydration of the cytoplasm and the production of special proteins that protect the spore's DNA. Figure 3.34 shows the major structures of an endospore.
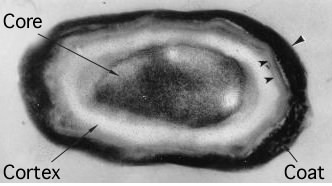
Figure 3.34. An endospore. An electron micrograph of an endospore of Bacillus subtilis showing the core, cortex and coat. (Source: M. Serrano, et al. 1999. J. Bacteriol. 181:3632-3643)
Endospores can be divided into several important parts (Fig 3.51). The center of the endospores contains the core, and it consists of the cytoplasm, DNA, ribosomes, enzymes, and everything needed to function once returned to the vegetative state. The core is dehydrated, which is essential for heat resistance, long-term dormancy, and maximum chemical resistance. Calcium dipicolinate is a major component of the core and plays a role in resistance to wet heat and UV light. The cortex surrounds the core and comprises two layers, a thin, more densely staining layer that is similar in structure to the vegetative cell wall and a thicker, less dense layer containing modified peptidoglycan. Two major modifications are present. First, there is less cross-linking with only 3% of the muramic acid present in the peptidoglycan of the cortex participating compared to 40% of muramic acid in the vegetative cell wall. Second, much of the muramic acid is modified to a muramic-β-lactam structure. Both of these modifications of the cortex appear to be important in germination. Muramic-β-lactam serves as a specific target for lytic enzymes that activate during germination, and the lower cross-linking enables easier outgrowth. Outside of the cortex is the spore coat composed of more than two dozen different types of proteins. There is some evidence that cross-links connect these proteins. This covalent connection between coat proteins probably contributes to the spores' resistance.
Some microbes make other types of resting structures
Endospores are not the only type of spore made by microorganisms. Azotobacterspecies and many others are known to form cysts, which are dormant cells with thickened cells walls (Figure 3.35). Cysts are resistant to desiccation and some chemicals, but cannot withstand high temperatures as endospores can.
The actinomycetes are a large group of spore-forming, gram-positive bacteria that grow by forming long tubules called filaments. Under nutrient-poor conditions, these filaments differentiate into round resting structures termed spores. In contrast to endospores and cysts, these structures are part of the reproductive process. The developmental process to create an actinomycete spore is less complex than that of the endospore. It involves the simple formation of cross walls that divide the filament into sections, each containing a chromosome. These then differentiate into mature spores. During this process, the bacterium lays down a tougher cell wall, and there is a conversion of the cytoplasm to a dormant state so that the spore becomes more resistant to heat and chemicals, though not as hardy as an endospore. Actinomycete spores can survive for long periods (for years) and germinate into vegetative cells when appropriate growth conditions are present. Many different genera are capable of forming this type of spore, and the ability to form these structures does not seem to correlate with any group of microorganisms.
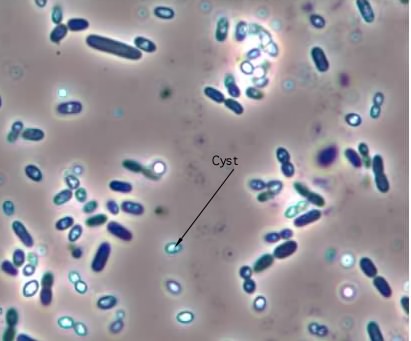
Figure 3.35. A cyst. The micrograph shows cysts and cells of Azotobacter vinelandii at 1000 X magnification. The cysts are the phase-bright objects, while the cells are darker.
Heterocysts are differentiated cells that specialize in nitrogen fixation
Some photosynthetic filamentous cyanobacteria are capable of forming specialized structures called heterocysts. These are rounded structures distributed at regular intervals along the string of vegetative cells, as shown in Figure 3.36 or at one end. Heterocysts evolved to solve the problem of performing plant-like photosynthesis (which produces oxygen) and at the same time fixing N2to ammonia (a process that involves enzymes that inactivate in the presence of oxygen). The single focus of the heterocysts is to fix N2, while the rest of the cells perform photosynthesis (and divide), thus keeping the two processes separate. Heterocysts develop a surface that is impermeable to gasses, and begin synthesizing large amounts of nitrogenase, the protein that fixes N2. Importantly, these heterocysts maintain some permeability to the cells on either side. Neighboring cells take N2from the atmosphere and pass it along to the heterocyst, which reduces it to NH3and returns fixed nitrogen to its neighbors. The neighboring cells also prevent oxygen from reaching the heterocyst. Heterocysts are essentially specialized organs for the "multi-cellular organism" represented by a chain of cyanobacterial cells that only form when nitrogen is limiting. The regulation of this developmental cycle is intriguing and serves as a simple example of multicellular development in a unicellular organism.
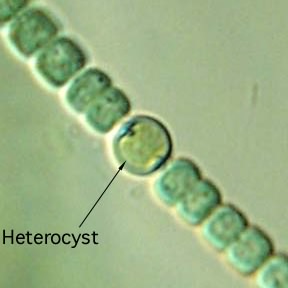
Figure 3.36. A heterocyst of a filamentous cyanobacteria from the Anabaena genus. Heterocysts are common in several different groups of cyanobacteria and are the site of nitrogen fixation. Note the slightly enlarged size and distinct shape of the heterocyst when compared to the vegetative cells on either side. (Source: Michael Clayton, University of Wisconsin-Madison)
This ends our survey of the cellular structure of bacteria. In the remaining sections, we look at the major structural differences that distinguish the archaea and eukaryotes from the bacteria.
Key Takeaways
- Cells can also differentiate to form visible structures that are not vegetative cells. Spores and cysts are resting structures that are a form of suspended animation.
- These structures allow a cell to avoid environmental challenges and germinate at a later time when conditions are better.
- Endospores can survive for very long periods and survive very harsh conditions.
- Heterocysts are another structure that cyanobacteria form to help them fix nitrogen. Heterocysts allow cyanobacteria to fix nitrogen, an anaerobic process, at the same time that they are performing photosynthesis, a process that generates oxygen.
3 - 8 Archaeal Structure
Learning Objectives
After reading this section, students will...
- Be able to explain the differences between the three domains of life.
- Be able to describe the unique properties of Archaea.
Before 1977, prokaryotes were phylogenetically organized into one group termed the Monera. The discovery of the extreme thermophile Thermus aquaticusin the hot springs of Yellowstone National Park by Tom Brock and the subsequent analysis by Carl Woese and others began a series of experiments that would change how scientists think about the organization of life on this planet. This revolution is the subject of later chapters, but here we will discuss the structural differences between this new group, the Archaea, and the Bacteria. "Bacteria" and "Archaea" refers to the formal phylogenetic classification of organisms, but we will typically refer to them more casually as groups of organisms and then use "bacteria" and "archaea". The third domain consists of eukaryotic organisms, both microbes and multi-cellular, and is referred to as Eukarya. As explained early in this chapter, it has become evident that eukaryotes arose when certain bacteria became engulfed in archaeal cells, eventually becoming organelles. Not surprisingly then, Archaea is a group of microbes that share some things in common with Bacteria, others with Eukarya, and still have other properties that are all their own. (see Table 2.5) In the following section, we highlight some of the major structural features of the Archaea.
Table 2.5 Comparison of properties between Archaea, Bacteria and Eukarya
| Property | Bacteria | Archaea | Eukarya |
| RNA polymerase | 4 proteins Rifampicin sensitive | 8-10 proteins Rifampicin resistant | 12 proteins (RNA pol II) Rifampicin resistant |
| Transcription start site | Variable often contains a -35 and -10 region | TATA box | TATA box |
| Starting amino acid | formylmethionine | methionine | methionine |
| Lipids | ester-linked | ether-linked | ester-linked |
| Cell wall composition | G+ peptidoglycan G- peptidoglycan and outer membrane |
pseudopeptidoglycan or S-layer of proteins, glycoproteins, or polysaccharides |
none or cellulose |
Archaea are a unique form of life, as different phylogenetically from bacteria as they are from eukaryotes.
The major differences between Archaea and other domains of life
RNA polymerase
DNA-dependent RNA polymerases in bacteria and archaea are different from each other. The primary RNA polymerase of all organisms is responsible for creating messenger RNA that the ribosome then translates into proteins. Bacterial RNA polymerase is relatively simple, containing five different proteins. In contrast, RNA polymerase from methanogens and halophiles (both Archaea) contains eight proteins. In hyperthermophiles (Archaea) RNA polymerase is even more complex, containing ten proteins. None of the archaeal RNA polymerases are affected by the antibiotic rifampicin, a known inhibitor of the bacterial RNA polymerase. The eukaryotic RNA polymerase responsible for most mRNA transcription (termed RNA polymerase II) contains 12 proteins and is similar to the RNA polymerase in the archaea.
RNA polymerases from all organisms recognize a variety of start sequences or promoters (Figure 3.37). A promoter for mRNA transcription in bacteria is recognized by the sigma protein and has two recognition zones about 10 and 35 bases before the transcription start site. In archaea and eukaryotes the transcription recognition sequence is a TATA sequence (termed the TATA box) and transcription is regulated by various protein transcription factors that bind to regions near the TATA box and then recruit RNA polymerase.
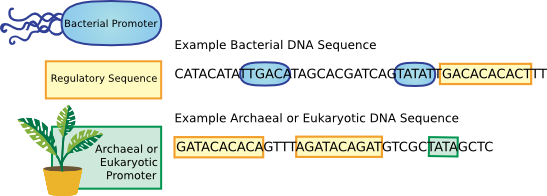
Figure 3.37. Control elements in bacteria and archaea. The best-studied bacteria have promoters at two positions upstream of the transcription start site that are recognized by the σ factors associated with RNA polymerase. The transcriptional control elements in the archaea are much more like those found in eukaryotes, where RNA polymerase is recruited to a promoter by interaction with protein regulatory factors, and transcription begins downstream of a TATA box.
Translation and ribosomes
The structural make-up of the ribosomes of bacteria and archaea are similar in many respects. The ribosomes of archaea and bacteria are of the same size (70S) and are smaller than those of eukaryotes (80S). However, most of the archaea's ribosomal proteins, translation factors, and tRNAs more closely resemble their counterparts in eukaryotes. Mixing experiments with ribosomes from E. coli(Bacteria), Sulfolobus(Archaea) and yeast (Eukarya) have shown functional substitution between Archaea and Eukarya. Specifically, mixing a small subunit from a yeast ribosome with the large subunit from Sulfolobus, results in a ribosome that is capable of translation in vitro. In contrast, mixing a small subunit from yeast with the large subunit of E. coli resulted in no translation. Also, in all organisms translation begins at a start codon (typically an AUG in the mRNA). In bacteria, this codon causes the insertion of formylmethionine, while in archaea and eukaryotes, it results in insertion of an unmodified methionine.
Lipids
Archaeal membranes have features unlike those found in either eukaryotes or bacteria (Figure 3.38). In eukaryotes and bacteria, the fatty acids are attached to the glycerol backbone by ester bonds. In archaea, ether linkages link the polar groups to their long-chain alkyl groups. Also, the stereochemistry of lipids from archaea is primarily of the S form, while that of bacteria and eukaryotes is of the R form. In some archaea, the hydrophobic chains attached to the glycerol backbone are twice the normal length and pass completely through the membrane, attaching to a second backbone on the opposite side. This linked membrane adds extra stability, and these dual lipids are often found in archaea living in extreme environments.
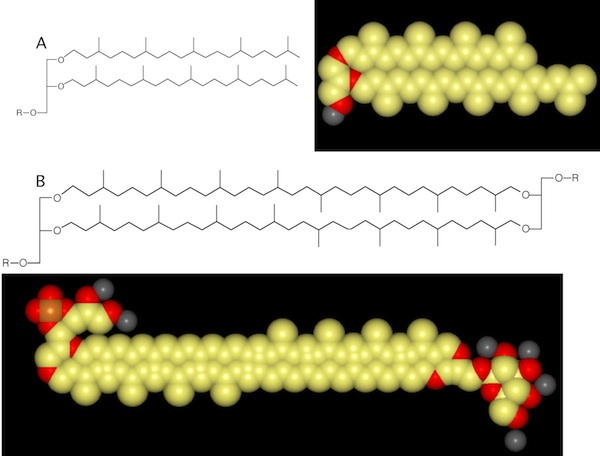
Figure 3.38. Ether-linked lipids. Archaea have an ether linkage between the long alkyl chain and the glycerol backbone, which is in contrast to the ester linkage seen in bacteria and eukaryotes. (A) An ether-linked lipid show as the chemical structure and a space-filling model. (B) A tetraether lipid, showing a general chemical structure. An example of a tetraether lipid from the thermophilic archaeal species Thermoplasma acidophiliumis shown as a space-filling model.
Cell walls
Most microorganisms in the Bacteria domain contain cell walls with peptidoglycan, with N-acetyl muramic acid being the signature molecule for the presence of peptidoglycan. Archaea are considerably more diverse in the composition of their cell walls. They lack peptidoglycan, but some contain pseudomurein that has a similar structure, as shown in Figure 3.39. N-acetylalosaminuronic acid replaces N-acetylmuramic acid in the molecule's backbone, and each glycan unit is linked together using 1,3 glycosidic bonds instead of the 1,4 glycosidic bonds seen in peptidoglycan. Many archaea do not contain peptidoglycan in any form, instead covering the outside of the membrane with proteins, glycoproteins, or polysaccharides. Scientists refer to these cell walls as S-layers. In any case, the function of the cell wall remains the same, containing the cytoplasm and giving the microbe its shape.
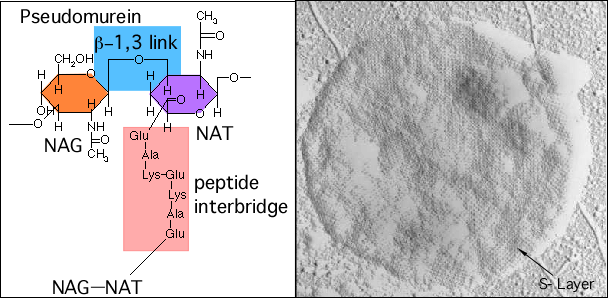
Figure 3.39. A comparison of cell wall structure in Archaea. Archaea have several different types of cell wall. Some contain a structure reminiscent of peptidoglycan called pseudomurein. The chemical formula is pictured on the left. Other microbes will have a surface layer (S-layer) composed of repeating units of one or a few proteins, glycoproteins or sugar. These crystal lattices serve to protect the cell. The micrograph on the right shows the surface of Methanospirillum hungateicells. Note the regular repetition of the pattern on the outside surface. (Source: M. Firtel, et al.1993. J. Bacteriol. 175:7550-7560)
These differences between archaea, bacteria, and eukaryotes indicate important differences between these life forms, and phylogenetic analysis bears out this conclusion. The archaea are more divergent from bacteria than we are from an amoeba. This significant divergence indicates that the common progenitor of these groups existed very early in evolution. Analysis and classification of the archaea has fundamentally changed how biologists think about organismal diversity.
Key Takeaways
- Archaea are a different life form, as different from bacteria as we are from E. coli.
- Archaea share traits that are similar to eukaryotes. The RNA polymerase of Archaea and the promoter structure is similar to eukaryotes.
- In addition, translation of proteins inserts a methionine as do eukaryotes, instead of a formylmethionine, as bacteria do.
- Archaea do have some unique traits, including ether-linked lipids and unique cell wall structures that can be formed from pseudopeptidoglycan or an S-layer.
3 - 9 Eukaryotic cells have much in common with prokaryotic cells
Learning Objectives
After reading this section, students will...
- Know that there are many basic similarities between eukaryotic and prokaryotic cell structures, but some structures are different.
- Be able to explain that eukaryotes are the only domain of life to have organelles and why that is true.
- Be able to explain the role of the highly organized cytoskeleton in Eukaryotes.
- Be able to explain how transport is different in eukaryotic cells in comparison to prokaryotes.
- Be able to explain how the structure and function of the nucleus affects transcription and translation.
- Be able to explain how the process of messenger RNA synthesis differs in Bacteria and Eukarya.
This section will focus on describing some of the differences between Bacteria and Eukarya. It will not be an exhaustive review of eukaryotic structure, as you have learned this material in your high school or college biology class.
After the recent journey through the bacterial cell, you may have started to wonder about your cells or other eukaryotic cells. How many properties do we share with bacteria? How are we different? It turns out, as you might expect, we share some basic things in common, but other structures are very different.
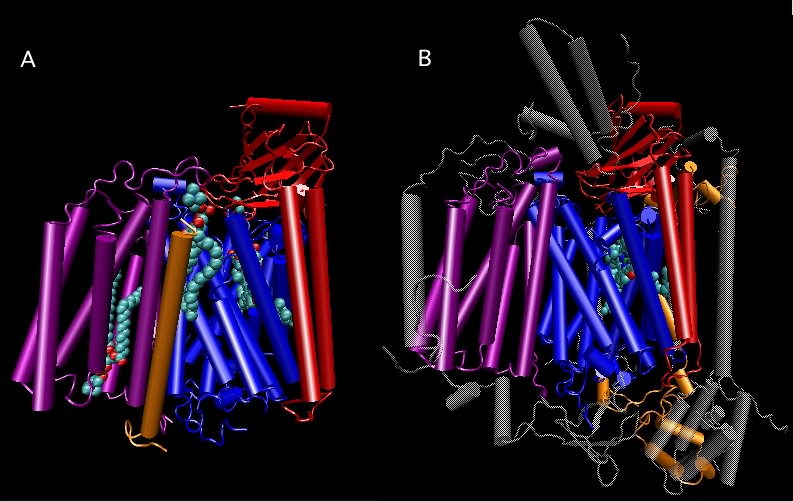
Figure 3.40. Comparison of cytochrome oxidase from bacteria and bovine.. Molecular models of cytochrome oxidase from Rhodobacter sphaeroides (A) and bovine (B) are compared. Each protein is a complex of several distinct proteins, but the four polypeptides shown in color have a high degree of similarity in both their sequence (see Figure 3.41) and structure. The structural similarity should be obvious in this view. Such structurally similarity cannot have arisen by chance, but must reflect the evolution of each from a single ancestor. The gray polypeptides in the bovine cytochrome oxidase are not found in the bacterial protein.
The basic building blocks of the cell, such as nucleic acids, amino acids and sugars are identical. Macromolecular organizations such as chromosomes and membranes have many similarities. Many proteins in eukaryotes, especially those that carry out essential cell functions, have homologs in bacteria that share a high degree of sequence and structural similarity. An example that illustrates this point is the respiratory enzyme cytochrome oxidase. As shown in Figures 3.40 and 3.41, a comparison of cytochrome oxidase from the bovine and Rhodobacter sphaeroides reveals a near-identical arrangement of the catalytic proteins and high sequence similarity. However, the cytochrome oxidase in the bovine has several other polypeptides that serve a structural role.

Figure 3.41. Sequence comparison of cytochrome oxidase from three species. A sequence comparison of cytochrome oxidase showing the high degree of identical amino acids between these very different species: cows and two different bacteria. The colored boxed indicate where the amino acids are identical or similar among the three sequences and the different colors refer to different classes of amino acids.
The DNA in all organisms is chemically similar but the organization of the helix into higher-order structures varies. Eukaryotes contain a larger number of histones (chromosome-binding proteins). In many cases, during cell division, compaction of the chromosome occurs so that they are visible using light microscopy.
The basic mechanism of converting genetic information into proteins is also rather similar. Most of the central components of this process show sequence similarities across biology and substantial functional similarities. The fact that eukaryotes have nuclei means that mRNA must be transported outside of that structure before translation can begin, but the process is otherwise rather similar.
The most conserved of all structures is probably the membrane. Membranes enclose all living systems and every membrane contains amphipathic lipids. The exact chemical structure of the lipids is often different depending upon the species and its environment, but the overall arrangement of the membrane is the same. Membranes perform remarkably similar functions in all species: keeping the cytoplasm in and the environment out. Cholesterol is common in the membranes of eukaryotes, but is uncommon in bacterial and archaeal membranes.
Things that are different between eukaryotes and prokaryotes
Eukaryotes are typically more complex than prokaryotes and appear more organized when examined in the microscope. This organization into different intracellular compartments likely reflects the demands of a more complex cellular system. It may also be that the formation of these separate organelles allowed the subsequent evolution of more elaborate cells. Figure 3.42 diagrams the various structures found in the typical eukaryotic cells. Only photosynthetic organisms contain chloroplasts. The organelles are the nucleus, the mitochondria, the endoplasmic reticulum and the Golgi apparatus. Internally eukaryotic cells have a cytoskeleton that determines cell structure, and plants supplement this with cell walls. We discuss these structures and their functions in eukaryotic cells in the following sections.
Figure 3.42. Eukaryotic cell structure. A diagram of the common structures found in eukaryotic cells.
Microfilaments and microtubules
The cytoskeleton is a network of filaments and fibers found in the cytoplasm of many eukaryotic cells. It serves four known roles in cells.
- The cytoskeleton helps to determine the shape of the cell.
- It is involved in cell division, allowing the separation of the chromosomes into daughter cells.
- The cytoskeleton moves organelles around the cell.
- It is involved in motility either through the use of flagella or by amoeboid movement.
The cytoskeleton components can be divided into three classes based upon the filaments' size, distribution, and function. Microfilaments, made from actin, are the smallest at 4 to 6 nm in diameter. These lie beneath the surface of the cell membrane and are anchored to it, forming a web-like structure. They dictate the cell's shape and can also be involved in motility by contraction or expansion of the filament. Filaments may also tether organelles to the membrane and help move them around the cell. This movement can be important for the modulation of organelle function. Intermediate filaments are 10 nm in diameter and made of keratin, the same protein found in hair and fingernails. These filaments take different forms and are present in many types of cells, but their exact function is unknown. They may play a structural role similar to some microfilaments. Figure 3.43 shows some examples of cytoskeletal elements
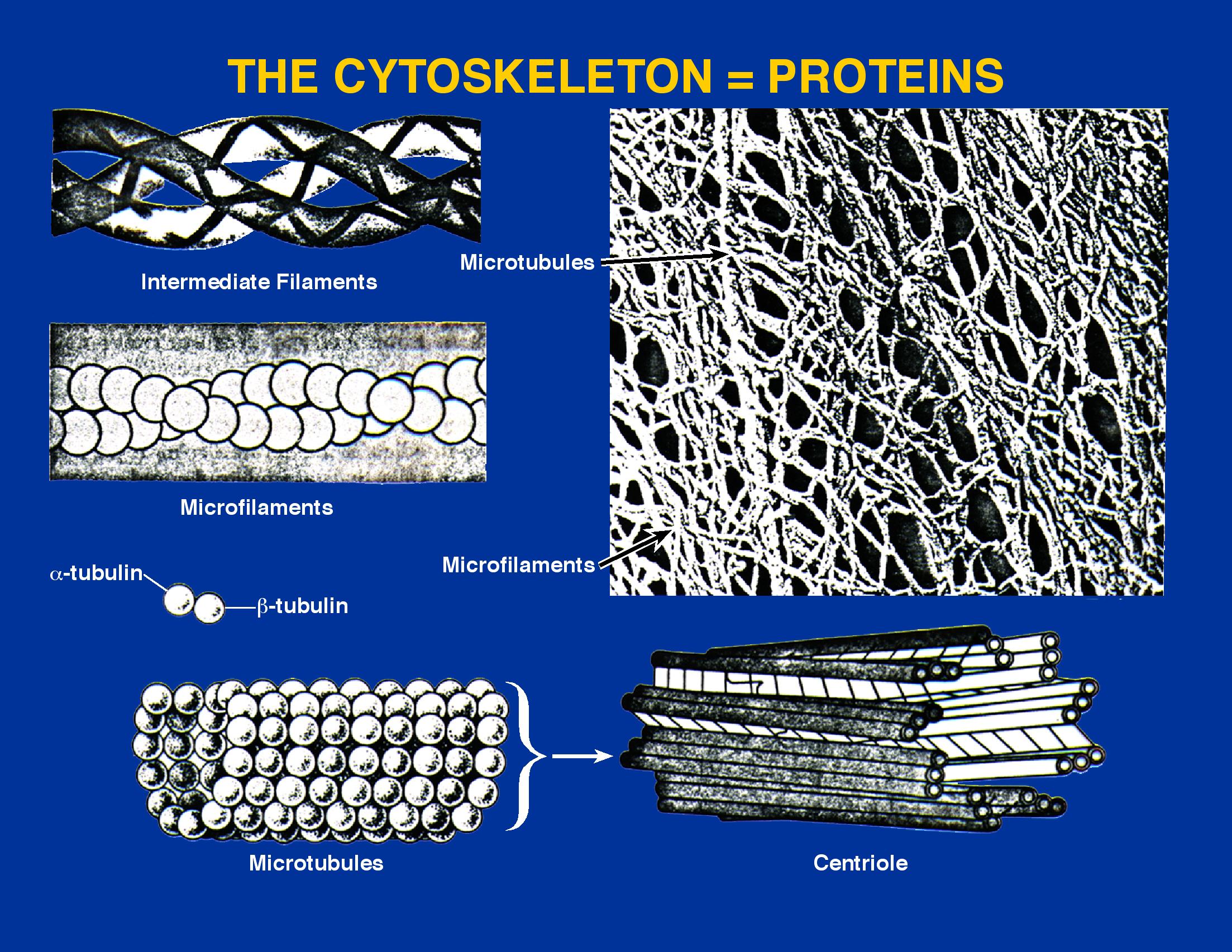
Figure 3.43. Cytoskeletal elements. Eukaryotic cells have several different types of scaffolding proteins to help them keep their shape. Microfilaments, microtubules, and centrioles are all important structural elements.
In addition to the above, there are more complex fibers and structures. Microtubules are hollow cylindrical structures 20-25 nm in diameter containing tubulin as the major structural protein. Tubulin polymerizes into a helical cylindrical structure, and thirteen of these protofilaments then combine to make a microtubule. Microtubules can form the basis of many different structures. Many of these structures form to perform a necessary function and are then disassemble. Often microtubules form centrioles that contain nine sets of microtubules arranged in a circular matrix. These are 4.0 µm long and 1.50 µm wide and are usually found in pairs at right angles to each other. Centrioles are important in proper chromosome segregation and cell division, as discussed in the section on the nucleus below. Microtubules are also part of the basal bodies of flagella and cilia.
Cilia and flagella are examples of more permanent structures that contain microtubules. Both cilia and flagella have a similar structure of nine pairs of microtubules arranged in a circular fashion around a tenth pair that runs down the center. Both are attached to the membrane and project into the environment. In a process that requires energy, the microtubules in the outer ring are moved with respect to each other, causing the cilia or flagella to bend and snap back in a whip-like fashion. This bending causes the movement of liquid near the structures such that spent liquid with few nutrients and waste products is moved away from the cell and replaced by fresh liquid containing nutrients and oxygen. The beating of cilia and flagella can also push the cell through its environment. Cilia are 2 to 10 µm long and 0.5 µm wide and are shorter and typically more numerous than flagella, with hundreds of them on some types of cells. An example of a ciliated organism is the unicellular protist Paramecium found in freshwater ponds. The microbe is a predator of bacteria, and motility is vital in this lifestyle, both for chasing down prey and moving away from danger. Cilia cover the surface of Parameciumand move the organism through the environment by beating in a coordinated fashion.
Flagella are 5-100 µm in length, and there are typically only one or two per cell. Eukaryotic flagella are larger than those found on bacteria or archaea and have a more complex structure. Flagella are found in many unicellular creatures, and their primary role is cell motility.
Endoplasmic reticulum
The endoplasmic reticulum (ER) is a finely divided system of interconnected membranes, consisting of tubules and vesicles that loop through the cell and are contiguous with the nuclear membrane. A drawing of the ER is shown in Figure 3.44. It functions in the synthesis of membranes and membrane proteins and is also involved in protein secretion. Not surprisingly, the ER is especially prominent in cells doing a large amount of protein secretion. The ER works very closely with the Golgi apparatus (see below) to carry out these functions. No structure in bacterial cells is analogous to the ER, but many of the same functions occur on the inside surface of the cellular membrane in bacteria. ER comes in two types: rough ER and smooth ER.
Figure 3.44. The endosplasmic reticulum. Eukaryotic cells contain a network of passages that connect various elements of the cell and are also important in secretion.
Rough ER gets its appearance from the presence of ribosomes on its surface, as seen in the electron microscope, and its function is the production, processing, and export of proteins. During translation, an appropriate signal guides the ribosome to the ER, causing protein synthesis directly across the membrane into the lumen. These proteins may be processed or modified by the addition of carbohydrates to form glycoproteins. After processing, proteins move slowly through the ER and are packaged into vesicles of ER membrane called transition vesicles. These release from the ends of the ER and move via the cytoskeleton to either to the Golgi apparatus or to the plasma membrane. Once the transition vesicle contacts the Golgi (or plasma membrane), the two fuse and release the contents of the vesicle into the target compartment.
Smooth ER does not contain ribosomes, and the lumen and membrane of smooth ER contain a variety of enzymes that perform many functions, including modification of toxins and synthesis of steroids.
Golgi apparatus
The Golgi apparatus is an organelle containing a double membrane, and it is mainly devoted to the processing of proteins synthesized in the ER. A drawing of the Golgi apparatus is shown in Figure 3.45. It is found in many eukaryotic cells, but it lacks a well-formed structure in many fungi and ciliate protozoa. It consists of regions of stacked contiguous membranes containing no ribosomes. Each membrane sac is 15 to 20 µm thick and separated from the next stack by about 30 µm. A complex network of tubes and vesicles extend from the edges of these sacs into the surrounding cytoplasm. The Golgi has a definite polarity with membranes near the ER (the cis face) having a different shape and enzyme content than those at the opposite end (the trans or maturing face). Studies of the Golgi apparatus appear to show material flowing into the cis face from vesicles through the apparatus and then exiting at the trans face.

Figure 3.45. The Golgi Apparatus. The Golgi apparatus is involved in the glycosylation and proteolytic processing of proteins that are to be secreted into various cellular organelles or to the outside environment. Proteins often pass through the Golgi apparatus as part of their maturation
The role of the Golgi apparatus is to package material for export, but its exact function varies depending upon the organism. For example, Giardia and Entamoebautilize Golgi apparatus to form cell walls during cyst formation. It often participates in the synthesis of cell membranes and the final processing of proteins before export. In many cases, the Golgi apparatus contains glycosylation enzymes that add sugars to proteins as they move through its lumen. The type of glycosylation that takes place is dependent upon signals contained within the protein sequence. The Golgi also processes enzymes using proteases, which clip at specific amino acids sequences to form mature proteins and hormones. These mature proteins then move to their final destinations, which may be in the membrane, in lysosomes or secreted into the environment.
Lysosomes
One of the most important functions of the Golgi apparatus is the synthesis of lysosomes, an organelle that is present in a variety of eukaryotic cells. Lysosomes are spherical structures enclosed in a single membrane that can vary in size from 50 nm to several µm. They are involved in intracellular digestion and contain enzymes (called hydrolases) that digest many types of macromolecules. Hydrolases function best under acidic conditions (pH 3.5 to 5.0) and the lysosome maintains this pH by membrane proteins that pump protons into its interior. Ribosomes synthesize enzymes bound for lysosomes on the ER. They then move through the smooth ER and Golgi apparatus before being package into lysosomes. In some cases, lysosomes can also bud off from the smooth ER.
Lysosomes serve a variety of functions depending upon the cell type. In unicellular eukaryotes, they are often digestive structures that take bacteria or other substances from the outside environment and degrade them into usable nutrients. In mammals, lysosomes serve to eliminate unwanted particles, either cell structures that are no longer needed or foreign macromolecules (from viruses or bacteria) that have invaded the cell. In any case, the hydrolytic enzymes and low pH typically inactivate and then degrade any particle that enters the lysosome
Eukaryotic cells absorb things by endocytosis
Lysosomes are an important part of endocytosis, a process that eukaryotic cells use to take up particles from the environment. Figure 3.46 illustrates endocytosis. The process creates membrane-bound cavities filled with fluid and solid materials. Larger membrane-enclosed cavities are called vacuoles, while smaller ones are called vesicles. Endocytosis comes in two forms, phagocytosis and pinocytosis. Phagocytosis involves the engulfment of large particles, even microorganisms, into membrane-bound compartments. It is a process used most often in the immune system, and the chapter on the innate immune system describes it in detail. Pinocytosis involves the recognition of specific particles in the environment, as described below. Unicellular eukaryotic microbes use the process to ingest food, while multicellular organisms use it to take in certain macromolecules traveling from other parts of the organism.
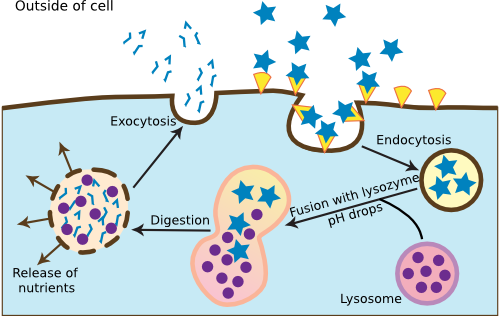
Figure 3.46. The process of endocytosis. Eukaryotic cells absorb materials from the outside by using endocytosis. This is a standard pathway through which most material enters the cell.
Pinocytosis begins when protein receptors on the cell surface bind to the target molecule to be ingested. The bound particle migrates to an area of the membrane that is rich in a protein called clathrin. This protein forms a matrix and causes an indentation in the membrane called a clathrin-coated pit. The entrance of a receptor:particle complex begins the invagination process at the pit that results in the engulfment of the incoming particle inside a membrane vesicle. This vesicle is called an endosome and once inside the cell, proteins in the endosome membrane begin to pump protons inside, dropping the internal pH of the endosome. The clathrin on the endosome membrane then migrates back to the plasma membrane to repeat the cycle. The endosome fuses with a lysosome to form an endolysosome, which causes the degradation of the ingested particle. The cell then transports the valuable breakdown products to their destinations. The spent endolysosome is called a residual body and sometimes fuses with the plasma membrane to release remaining compounds into the environment.
The ER, Golgi apparatus, lysosomes, and endosomes seem to operate as a coordinated whole, functioning to import and export materials. It is probably correct to think of these structures as a functional unit, and the term vacuome has been coined to describe them. Ribosomes in the ER manufacture proteins and the ER and Golgi modify them. The mature proteins eventually find their way to the plasma membrane for export or become part of lysosomes. Lysosomes serve in the endocytotic pathway to take up materials and process them for cell use. They also digest spent cell constituents. All of these processes occur within membrane structures. This carefully controls the import and export of materials from the cell. Considering the extent of these structures in the cell, it is remarkable that the membranes of these structures, especially lysosomes, never rupture. If they did, it would be catastrophic to the cell and would rapidly lead to cell death.
The nucleus holds the cell's genetic material in eukaryotes
While the bacterial cell does seem to sequester its chromosome to a portion of the cytoplasm, no demarcation divides the nucleoid from the rest of the cell. In eukaryotes, the nuclear membrane separates the cell's DNA from the cytoplasm. The nucleus is the largest and most visible organelle of eukaryotic cells. It contains almost all the cell's DNA and is the site of chromosome replication and transcription. It has two layers of membrane encircling it called the nuclear envelope, with the outer layer being contiguous with the ER. Scattered throughout this nuclear envelope are circular openings known as nuclear pores. These pores are highly discriminatory, allowing easy movement in and out of the nucleus of only appropriate macromolecules such as proteins with specific sequences.
In eukaryotes, the chromosome is not a single circular piece of DNA like most prokaryotes. Rather, most eukaryotic cells split their genomes into several linear chromosomes, with each cell containing at least two copies of each chromosome. The exceptions are those cells specializing in reproduction and only contain one copy of the cell's chromosomes. Each piece of DNA is complexed with special basic structural proteins called histones that seem to be important in keeping the DNA organized. Proteins bind the DNA, and the entire set of DNA and associated proteins is called chromatin. For much of the cell cycle, chromatin consists of long DNA strands formed into beads by the association of histones along its length. Figure 3.47 shows a nucleus during mitosis, with the chromosomes visible.
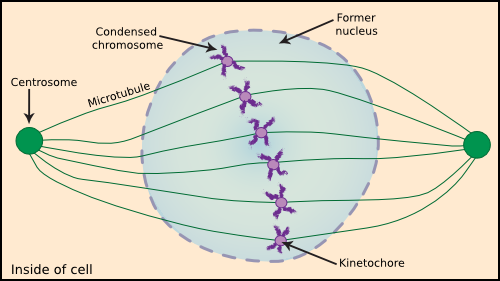
Figure 3.47. A eukaryotic cell in the middle of mitosis. Eukaryotic DNA replication takes place during the cell phase called mitosis. At this time, protein synthesis is halted and the chromosomes condense. The sister chromatids meet at the middle of the cell and then migrate to two separate poles. This movement is coordinated by centrosomes, kinetochores and microtubules.
In actively growing cells, the DNA is replicated from numerous sites rather than the single bi-directional origin in prokaryotes. These multiple DNA origins are necessary due to the much larger amount of DNA found in most eukaryotic cells. During division in prokaryotes, the cell appears to simply split in two, with each daughter cell receiving a chromosome. In contrast, eukaryotic cells go through a morphologically distinct phase, mitosis, to separate the chromosomes. One of the more important events of mitosis is the binding of additional histones and the contraction of the chromatin into compact structures that were called chromosomes due to their staining properties. (The original meaning of the term chromosomes is a colored body, but is now synonymous with a cell's DNA). The two daughter chromosomes formed during replication are physically separated into the separate daughter cells by the filaments called microtubules. These attach at one end to the chromosome at a region termed the kinetochores, and at the other end, they attach to one of two regions of the cell called a centrosome. By depolymerizing the microtubules at each centrosome, each daughter chromosome is pulled away from its partner and toward a region that eventually reforms as a new nucleus.
There are also some important differences in transcription between eukaryotes and prokaryotes. In eukaryotes, mRNA transcription takes place in the nucleus, and the finished mRNA moves through the nuclear pores and into the cytoplasm for translation by the ribosomes. The genes of eukaryotic cells also contain regions of largely unimportant DNA, termed introns, that do not code for protein. After the transcription of a gene into mRNA, enzymes remove these introns before translation. One set of nuclear proteins removes these sequences and splices the actual coding sequences (exons) together to make the finished mRNA. The finished mRNA then travels from the nucleus to ribosomes in the cytoplasm. Modifications of both ends stabilize the mRNA of eukaryotic cells. At the front end is usually a 5'-cap made of 7-methylguanine attached to the mRNA by a triphosphate linkage. At the 3' end of the mRNA is a long stretch of A bases (a poly-A tail) that have a role in mRNA stability, as mentioned earlier in this chapter. Finally, while it is quite common in bacteria to have several genes on each mRNA transcript, the vast majority of mRNAs in eukaryotes code for only a single protein product. Figure 3.48 shows the steps in gene expression in eukaryotes.
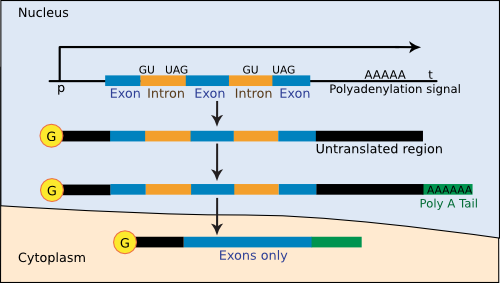
Figure 3.48. The steps of gene expression in eukaryotes. Processing genetic information into protein is more complex in eukaryotes than in prokaryotes. In part this is due to the existence of introns in most genes of "higher eukaryotes" (though introns are rare in yeast). These have to be removed, a poly-A tail added to the 3' end and a cap added to the 5' end of the mRNA before it reaches the cytoplasm for translation.
Eukaryotic cells also contain one or more dark-staining structures within the nucleus called nucleoli. Although they are not enclosed within a separate membrane, the nucleoli are complexes with separate granular and fibrillar regions. They are present in non-dividing cells but frequently disappear during mitosis and reappear after cell division. The nucleolus is the initial site of ribosome synthesis. This structure contains the DNA that codes for the ribosomal RNA genes. The ribosomal RNAs are synthesized and then processed to form the final rRNA molecules. These then combine with several ribosomal proteins synthesized in the cytoplasm to form an initial ribosomal complex. The entire complex then migrates out of the nucleus into the cytoplasm, where it combines with other proteins to form a ribosome.
Mitochondria and plastids are organelles of energy generation in eukaryotic cells
Mitochondria are involved in energy generation through respiration. Mitochondria have no fixed shape but often look like short rods in transmission electron micrographs when viewed along their long axis (Figure 3.49). Each mitochondrion contains two membranes. The outer membrane is smooth and serves as a selective barrier. The inner membrane is highly convoluted and folded and contains a high number of membrane complexes. Nutrients are oxidized inside the mitochondria by catabolic enzymes, and the high-energy electrons extracted are donated to a respiratory chain in the inner membrane. These enzymes then create a proton gradient, and this gradient is then used to synthesize ATP. ATP leaves the mitochondria, and it serves as a source of energy for the rest of the cell's machinery.
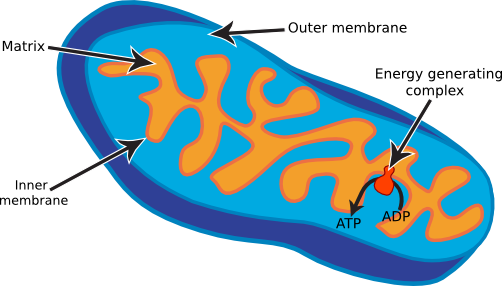
Figure 3.49. Mitochondria structure. Mitochondria are rod-shaped structures that resemble the shape of common bacteria. They contain two membranes, similar to what is found in gram-negative bacteria, and 70S ribosomes. Energy generation occurs at the inner membrane.
Plastids are specialized organelles involved in metabolism that are unique to plants and come in several forms. Amyloplasts are starch storage containers found in some plants. Chloroplasts are oval-shaped structures inside plant and algal cells containing an outer and inner membrane, as shown in Figure 3.50. The outer membrane serves a similar function to the outer membrane of mitochondria, while the inner membrane consists of a network of stacks of membranous disks, called thylakoids, which are attached together by narrow tubes of membrane. The thylakoid membranes are the centers of photosynthesis in eukaryotes. They contain enzyme complexes that capture light and produce ATP and high-energy electrons. These then power the synthesis of organic compounds from carbon dioxide.
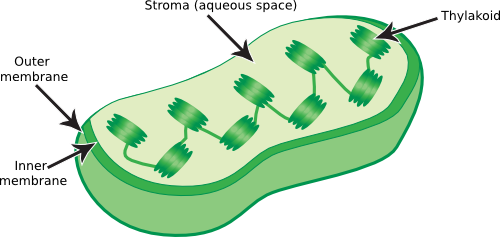
Figure 3.50. Chloroplast structure. Chloroplasts are the site of the light reactions of photosynthesis. Light striking the chloroplast is converted into a proton motive force and this is used to generate energy. Chloroplasts contain two membranes, again similar to gram-negative bacteria, and the chloroplast itself is a relative of cyanobacteria.
Mitochondria and chloroplasts each have a single circular chromosome and large numbers of ribosomes that are of bacterial (70S) and not eukaryotic (80S) form. The presence of two lipid bilayers, a circular chromosome, and 70S ribosomes is consistent with the evolutionary hypothesis that we have already explained near the beginning of this chapter.
The implications of eukaryotic structures on their growth
Eukaryotic cells are generally much larger than prokaryotic ones, and this difference in volume has several implications. First bigger cells can afford to have more things stored in the cytoplasm. This larger size means it is not as costly for a eukaryotic cell to have structures taking up space. In a prokaryote, space is at a premium, and the cell rapidly degrades anything not being used. This larger size may be one reason that organelles are possible. Second, larger cells have a lower surface-to-volume ratio than smaller cells, and therefore, prokaryotes effectively have more contact with their environment. This greater exposure can mean a more rapid response to changing environmental conditions. Finally, bigger cells have more of a challenge moving molecules within themselves. Prokaryotes can often depend on simple diffusion to move molecules around the cell, but this process might be too slow and inefficient in much larger cells. Eukaryotes overcome this by having specific transport mechanisms (i.e., microtubules) inside the cell.
Size constrains eukaryotes in two important ways: how fast they grow and what environments they can tolerate. The compartmentalization of the genome inside the nucleus limits the rate at which eukaryotic cells can divide. The complete cell division cycle in a multicellular eukaryotic organism depends on the cell type, but it takes at least 8 hours even in rapidly dividing skin cells. In unicellular yeast cultures, the shortest cell cycle is about 1.7 hours under ideal conditions. Due to their smaller genomes, lack of a nucleus, and the ability to couple transcription and translation, bacteria can grow much faster. Clostridium perfringens has been shown to go through a complete division cycle in at little as 6.6 minutes at 43 °C in beef cubes!
There are environments that have one or more physical characteristics that prevent the growth of most organisms. These extreme environments might be too hot, cold, acidic or alkaline for typical organisms to grow. However, a small subset of prokaryotes has evolved to take advantage of these environments and thrive, and prokaryotes always define the extremes of where life can exist. In part, this is probably because simpler cells have fewer "body parts" that must be changed for growth under very different conditions. In eukaryotes, they would need to modify their cytoplasmic contents to tolerate the extreme environment and the makeup of their organelles.
Key Takeaways
- At the molecular level, Bacteria and humans are more alike than different. However, these differences are important.
- First eukaryotes are larger, and this allows them to carry more DNA and other structures. The DNA can therefore code for more genes, enabling the control and regulation that is necessary in a multicellular organism.
- This complexity is manifested in the various visible structures of the cell, including: mitochondria (energy generation), the nucleus (DNA storage and transcription), chloroplasts (photosynthesis), the Golgi apparatus, the endoplasmic reticulum, lysosomes (the last three involved in protein export).
- The larger size of Eukarya cells necessitates the presence of a larger cytoskeleton, consisting of microfilaments and microtubules, involved in the transport of materials about the cell and the formation of structural components.
- The increased complexity of eukaryotes also makes eukaryotic cells less adaptable and unable to survive in extreme environments. Their large size slows the production of proteins, and eukaryotes grow more slowly than prokaryotes.
3 - 10 Instructions for buying Through the Microscope
If you enjoyed these first chapters on the history and significance of microbiology and on the chemical structure of organisms, consider buying the book. We designed Through the Microscope as an online resource, unlike most textbooks on the market that create online resources as an afterthought to sell you their overpriced hard copy. Thus, the pricing model is different. You are paying for a subscription to gain access to the material. It's only $30 and has stayed that way for over 25 years.
- First, register for the site using the registration form.
- After filling out your registration, you will get an Email from the site asking you to verify that you genuinely did register for the site. This email can end up in your junk folder, so please look there if you don't find it in your inbox.
- Follow the directions in the verification email and then log into the site.
- Once logged in, click the Buy Now button for Through the Microscope on the front page and follow the prompts. Once you are finished, please return to this site.
- You may need to reload the Table of Contents to gain access to any pages.
- If you have any problems with this process. Please contact the author using the contact form.
3 - 11 Summary
We have moved from within prokaryotic and eukaryotic cells to the outside, and this should provide a framework for thinking about the make-up of microbes. Some ideas that you should take away from this chapter are:
- While not obvious in the microscope, the prokaryotic cell is organized into functional units: (i) The nucleoid organizes, manages, and replicates the DNA. (ii) The cytoplasm carries out many of the reactions of the cell including biosynthesis and catabolism. (iii) The ribosomes are the site of protein synthesis. (iv) The membrane defines the borders of the cell and serves as a barrier to most molecules. (v) The cell wall defines a microbes shape and helps protect it from the environment.
- The building blocks that make up the average prokaryote are also found in all living things on this planet. We share more things in common with microbes than there are features that distinguish us.
- Although prokaryotes are small, they have a huge amount of molecular complexity. In a sense, multicellular organisms deal with different problems by forming different cell types, but prokaryotes must adapt each cell to specific environmental conditions. They are stunningly clever little machines.
In subsequent chapters, we will learn what all these various parts are doing.
If you enjoyed this free chapter on the structure of microbes, consider buying the book, for only $30.