Contact Us
Latest News
- Clues beginning to emerge on asymtomatic SARS-CoV-2 infection
- Back in November of 2020, during the first wave of the COVID-19 pandemic, I was teaching an in-person microbiology laboratory. One of my students had just been home to see his parents, and they all c…
- Read more
- Could there maybe be better uses of genetics and probiotics?
- Professor Meng Dong and his laboratory have created a probiotic that can metabolize alcohol quickly and maybe prevent some of the adverse effects of alcohol consumption. The scientists cloned a highl…
- Read more
- ChatGPT is not the end of essays in education
- The takeover of AI is upon us! AI can now take all our jobs, is the click-bait premise you hear from the news. While I cannot predict the future, I am dubious that AI will play such a dubious role in…
- Read more
- Fighting infections with infections
- Multi-drug-resistant bacterial infections are becoming more of an issue, with 1.2 million people dying of previously treatable bacterial infections. Scientists are frantically searching for new metho…
- Read more
- A tale of two colleges
- COVID-19 at the University of Wisconsin this fall has been pretty much a non-issue. While we are wearing masks, full in-person teaching is happening on campus. Bars, restaurants, and all other busine…
- Read more
Chapter 1 - The Relevance and History of Microbiology
1 - 1 Dedication
Writing a textbook is never an individual effort, and I would like to take a moment to thank all the people who helped along the way. Through the Microscope has been a labor of love for over a decade. Many people gave valuable input and made the way easier in many ways.
To the Bacteriology Department. Thank you for providing the opportunity and creating an environment where teaching and learning matter. It is an honor to work with such gifted and talented people—specifically, thanks to Robin, Jon, Diana, Jorge, Jo, Sarah, Melissa, Michelle, and Judy. I have learned much about learning, teaching, and life from each of you. And a special thank you to my former co-author, Gary Roberts, who has since retired from academic life and writing this book. As if you didn't have enough to do, you joined me in this quixotic adventure. Your contributions to the book have made it infinitely better than what I could have created by myself.
To my mother. Though you have been gone for over 30 years, your love and lessons stay with me. You taught me to work hard and instilled in me a passionate curiosity about the world. Your example and love still sustain me. To my father. You have always been someone I looked up to for how you moved through life, constantly overcoming the obstacles in front of you with a positive outlook. You were up to any challenges placed in front of you, and you met them all. I will never forget your smile and the light in your eyes.
To my children. For those times when dad was working on the book, and you needed something, I don't regret stopping to help. For the laughter, the sledding, the trips, and for just hanging out. I thank you. Your presence and sense of fun make life worth living.
Finally, to my wife. Thank you for your patience, as I spent many nights at the computer, instead of with you. For your sage advice, when I came up against a difficult problem. For your encouragement when I feared I would never get it done. And for your love, that makes everything I do worthwhile.
To all the friends who gave me advice and encouragement on the book, I owe you one, especially those of you who read preliminary copies and gave me advice. I cannot say how much it meant to me to have your guidance, your excitement, and counsel.
1 - 2 Introduction
This book will focus on the biology of small things, but what is microbiology? Microbiology could be defined as the study of organisms too small to be seen with the naked eye. Figure 1.1 shows the relative size of microbes compared to other living things. However, the relatively recent discovery of bacteria of near 1 mm in size has made this definition somewhat inaccurate. In the grand tradition of science, a new definition is in order.
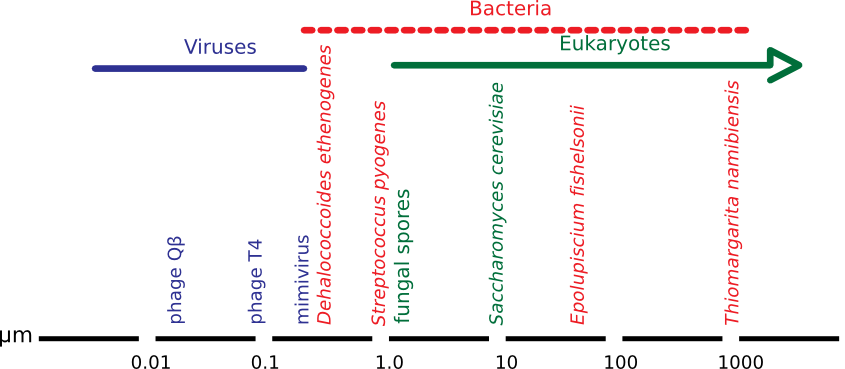
Figure 1.1. The relative size of microbes. Though microbes are small, they nevertheless span a large range of sizes from the smallest bacterial cells at ~0.15 µm to giant bacteria larger than 700 µm. The viruses depicted at the far left of the scale are even smaller.
We will consider microbiology to be the study of organisms that can exist as single cells, contain a nucleic acid genome for at least some part of their life cycle, and are capable of replicating that genome. This broad description encompasses an understandably large group of organisms, including fungi, algae, protozoa, and bacteria. Figure 1.2 has some examples of each. This definition would also include viruses, which microbiology texts traditionally discuss along with living organisms.
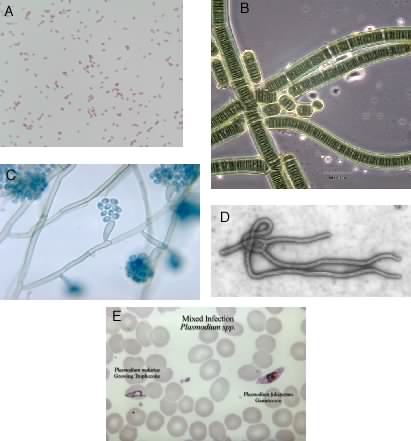
Figure 1.2. Some examples of the types of microbes present in the environment. Many different organisms fall under the definition of microorganisms. Shown here are: A, the bacterium Escherichia coli; B, a photosynthetic cyanobacterium; C, a fungus; D, Ebola virus; E, the protozoan malaria parasite. (Sources: B, Mike Clayton; C-E, CDC). Note that the scale on each of these pictures is different.
Microbiology also involves a collection of techniques to study and manipulate these small creatures. Because of their size, scientists use special instruments and methods to perform interpretable experiments on microorganisms. These methods are not restricted to microbes alone but have also found utility in working with populations of cells from higher organisms.
Microorganisms are everywhere, but why is it worth learning about them? The short answer is that they affect your life in many different ways. Before we begin our study of these creatures, we will first take a tour of some of their habitats and point out why your existence depends upon them. We will then briefly explore the history of microbiology.
1 - 3 Microbes have a large impact on human health
Learning Objectives
After reading this section, students will be able to...
-
Give some examples of the diseases microbes cause.
-
Explain the impact of vaccines, antibiotics and other advances on infectious disease in the developed world.
-
List some of the new illnesses caused by microorganisms that are emerging and the known pathogens that are becoming resistant to treatment.
-
Give examples of the mutualistic relationships that microbes form with other organisms.
If you ask the average person how microbes (or germs) impact their lives, they would immediately think of disease. This response is not a silly view, as pathogenic microorganisms have greatly affected human populations throughout our existence. Until about 1930, microbes were the major cause of death in humans. In 1900, infant mortality rates in some urban areas were up to 30%. From today's perspective, this is a horrendous statistic; over half of all infants did not make it to adulthood! With the advent of antibiotics, vaccines and better water sanitation, humanity has reduced the impact of pathogenic microbes, but they will always remain an important social concern. The discipline of microbiology emerged from the study of these diseases, and most advances in treating various ailments had their roots in this relatively young science. Figure 1.3 shows some important pathogens.
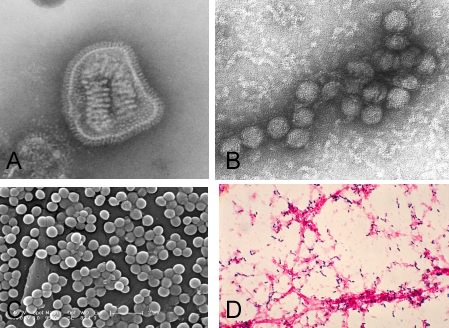
Figure 1.3. Some important pathogens. Many microbes cause disease in humans. Depicted here are several pathogens that cause important illnesses. A, Influenza virus; B, West Nile Virus; C, Staphylococcus aureus; D, Streptococcus pneumoniae. (Sources: A, Dr. Erskine/L. Palmer/ Dr. M. L. Martin; B, Cynthia Goldsmith; C, Janice Carr/ Jeff Hageman, M.H.S.; D, Dr. Mike Miller; all individuals are at the CDC.)
While death from infectious disease in the U.S. has been dramatically diminished, infection rates in developing nations remain unacceptably high. "Ancient" diseases continue to be a problem where nutrition and sanitation are poor, and emerging diseases such as Acquired Immunodeficiency Syndrome (AIDS) are even more dangerous for such populations. The Centers for Disease Control and Prevention (the U.S. government agency charged with protecting human health and safety) estimate that HIV infects about 9% of adults aged 18-49 in Sub-Saharan Africa. Yet as you can see in Table 1, AIDS is only one of a number of new diseases that have emerged. Many new diseases are viral in nature, making them notoriously difficult to treat, and they have no known cure. In addition, influenza and pneumonia are leading killers of the elderly, even in the U.S. and other developed nations. Even the common cold causes illness and misery for almost everyone and drains the productivity of all nations.
Disease due to food-borne pathogens is an increasing problem, primarily because of the consumption of improperly processed or stored foods. Understanding the sources of contamination and developing ways to limit the growth of pathogens in food is the job of food microbiologists.
Table 1.1 Disease-causing microbes and infectious diseases recognized since 1973.
| Year | Microbe/disease | Type | Health problem |
| 1973 | Rotavirus | Virus | Major cause of infantile diarrhea worldwide |
| 1975 | Parvovirus B19 | Virus | Severe anemia |
| 1976 | Cryptosporidium parvum | Parasite | Acute and chronic diarrhea |
| 1977 | Ebola | Virus | Ebola hemorrhagic fever/uncontrolled bleeding and kidney failure |
| 1977 | Legionella pneumophila | Bacteria | Legionnaire's disease |
| 1977 | Hanta virus | Virus | Hemorrhagic fever |
| 1977 | Campylobacter jejuni | Bacteria | Short-term diarrhea |
| 1980 | Human T-lymphotropic virus I (HTLV-I) | Virus | T-cell lymphoma-leukemial cancer of the blood |
| 1981 | Toxic strains of Staphyloccus aureus | Bacteria | Toxic shock syndrome |
| 1982 | Escherichia coli O157:H7 | Bacteria | Hemorrhagic colitis; hemolytic uremic syndrome |
| 1982 | HTLV-II | Virus | Hairy cell leukemia |
| 1982 | Borrelia burgdorferi | Bacteria | Lyme disease |
| 1983 | Human immunodeficiency virus (HIV) | Virus | Acquired immune deficiency syndrome (AIDS) |
| 1983 | Helicobacter pylori | Bacteria | Peptic ulcer disease |
| 1985 | Entercytozoon bieneusi | Parasite | Persistent diarrhea |
| 1986 | Cyclospora cayetanensis | Parasite | Persistent diarrhea |
| 1988 | Human herpesvirus-6 (HHV-6) | Virus | Roseola subitum/skin rash |
| 1988 | Hepatitis E | Virus | Liver infection; epidemic hepatitis |
| 1989 | Ehrlichia chaffeensis | Bacteria | Human ehrlichiosis/influenza-like infection |
| 1989 | Hepatitis C | Virus | Chronic liver infection |
| 1991 | Guanarito virus | Virus | Venezuelan hemorrhagic fever |
| 1991 | Encephalitozoon hellem | Parasite | Conjunctivitis |
| 1991 | New species of Babesia | Parasite | Atypical babesiosis/infection with fever, chills and fatigue |
| 1992 | Bartonella henselae | Bacteria | Catch scratch disease/bacillary angionmatosis |
| 1993 | sin nombre virus | Virus | Adult respiratory distress syndrome |
| 1993 | Encephalitozoon cuniculi | Virus | Infection with fever, chills and fatigue |
| 1994 | Sabia virus | Virus | Brazilian hemorrhagic fever |
| 1995 | HHV-8 | Virus | Associated with Kaposi sarcoma in AIDS patients |
| 2019 | SARS-CoV-2 | Virus | The cause of COVID-19 |
Source: WHO, The World Health Report 1996: 112
New infections continually appear. Having an available food source to grow on (humans) inevitably results in a microorganism that will take advantage. Some of these feeders will interfere with our well-being, causing disease.
Surprisingly, many diseases that have behavioral or genetic components we now suspect have microbial components. The clearest case is ulcers, which were long thought to be caused by stress and poor diet. However, the causative agent is a bacterium, Helicobacter pylori, and antibiotics can cure many of them. Work on other non-infectious diseases such as heart disease, stroke, and some autoimmune diseases also suggest a microbial component that triggers the illness.
Finally, some pathogenic microbes that had been "controlled" through the use of antibiotics are beginning to develop drug resistance and therefore reemerge as serious threats in the industrialized world as well as developing nations. Tuberculosis is an illness that was on the decline until the middle 80's. It has recently become more of a problem, partly due to drug resistance and partly due to a higher population of immunosuppressed individuals from the AIDS epidemic. Staphylococcus aureus strains are emerging that are resistant to many of the antibiotics that were previously effective against them. These staph infections are of great concern in hospital settings around the world. Understanding both familiar killers and new pathogens will require an understanding of their biology, and thus an understanding of the field of microbiology.
Microbes are often helpful, not harmful
From the beginning of microbiology, significant resources have been spent to understand and fight disease-causing microorganisms. You may be surprised to learn that only a small fraction of microbes are involved in disease; many other microbes enhance our well-being. The harmless microbes that live in our intestines and on our skin help us fight off disease. They actively antagonize other bacteria and take up space, preventing potential pathogens from gaining a foothold on our bodies. The microbial community in humans not only protects us from disease, but also provides needed vitamins, such as B12. We have entire communities of microorganisms in our digestive systems that contribute to our overall health. In fact, like all other large organisms, humans are consortia of different organisms - there are more non-human cells in and on our bodies than there are human cells! Recent experiments that have examined microorganisms inside our digestive tract by intensive sequencing experiments have revealed many interesting findings. More than 80% of the microbes in our guts have not been cultured. In addition, the microbiota of a person is unique to that person, and there are differences based upon body type and genetic background. These differences in our microbial partners have profound effects on the physical health of the individual.
Human health and nutrition also depend on healthy farm animals. Cows, sheep and other ruminant animals utilize their microbial associates to degrade plant material into useful nutrients. Figure 1.4 shows the cow, one example of a ruminant animal. Without these bacteria inside ruminants, growth on plant material would be impossible.

Figure 1.4. The cow as an example of a ruminant animal. In contrast to humans, ruminant animals have a complex system of stomachs that harbor large numbers of microorganisms. These microbes degrade the tough plant material eaten by the animal into usable nutrients. Without the assistance of the microbes, ruminant animals would not be able to digest the food they eat. (Source: Keith Weller, USDA.)
Commercial crops are also central to human prosperity, and much of agriculture depends upon the activities of microbes. For example, an entire group of plants, legumes, forms a cooperative relationship with certain bacteria. These bacteria convert nitrogen gas to ammonia for the plant, an important nutrient that is often limiting in the environment. Figure 1.5 shows a leguminous plant and the special structures on the roots that house these helpful bacteria. Most plants also form cooperative relationships with fungi, the mycorrhizal fungi, that help the plant sequester and absorb needed nutrients. Microbes also serve as small factories, producing valuable products such as cheese, yogurt, beer, wine, organic acids, and many other items. In conclusion, while it is less apparent to us, the positive role of microbes in human health is at least as important as the negative impact of pathogens.
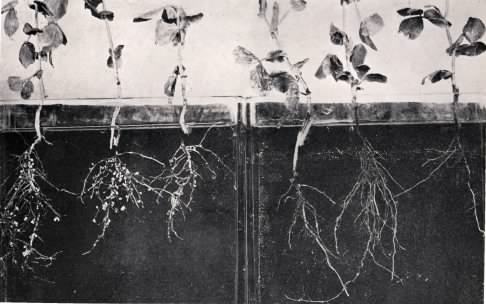

Figure 1.5. A leguminous plants.. Nitrogen-fixing bacteria form special structures, called nodules, on the roots of leguminous plants. In the picture, peas are shown that have either been exposed to bacteria (left) or not (right). The small bumps on the roots on the left are the nodules and contain millions of bacteria actively fixing N2 for the plant's use. Though not obvious from this figure, the plants on the left are more robust because of that nitrogen. (Source: E. B. Fred, et al., 1921. Soil Science 11:479-491.)
Key Takeaways
- Microbes cause many infectious diseases.
- Vaccines, antibiotics, and many other advances have lessened the impact of infectious disease in the developed world, but infectious disease in developing countries is high.
- New illnesses caused by microorganisms continue to emerge and known pathogens are becoming resistant to treatment.
- Microbes form important mutualistic relationships with all sorts of organisms.
- Many of these relationships are important from a human perspective.
1 - 4 Microbes have profound effects on the environment
Learning Objectives
After reading this section, students will be able to...
- Explain how cyanobacteria and algae in the oceans are responsible for most photosynthesis and are a major sink for carbon dioxide, a greenhouse gas.
- Describe the role that microbes play in the recycling of nutrients in the environment.
- Compare the role microbes play in energy production, both good and bad.
- Describe how microbes are used as model organisms.
- Explain how the tractability of microbes has made them essential in understanding core questions in biology.
- Describe how microbes are classified and what this means for all life on earth.
- Explain the roles microbes play in molecular biology.
Whether measured by the number of organisms or by total mass, the vast majority of life on this planet is microscopic. These teaming multitudes profoundly influence the make-up and character of the environment in which we live. It may surprise you that we know very little about the microbes that live in the world around us because less than 2 % of them can be grown in the laboratory. Understanding which microbes are in each ecological niche and what they are doing there is critical for our understanding of the world. Figure 1.6 shows some examples of environments where microbes are present.
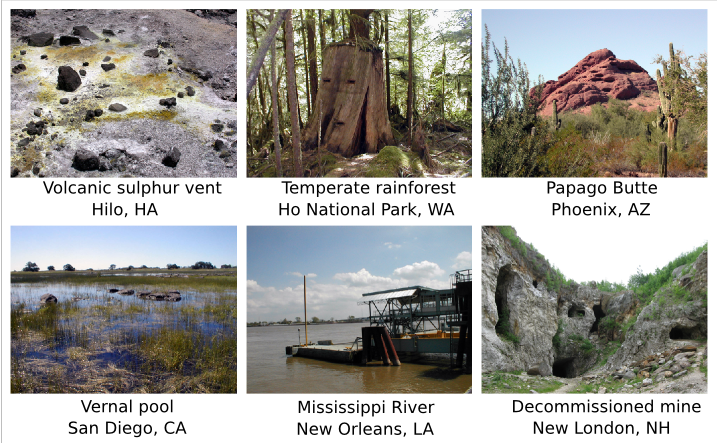

Figure 1.6. A sampling of the different types of environments where microbes flourish. Microbes are capable of growing in a wide variety of environments. Bacteria will grow in frigid glaciers to boiling volcanic springs, dry sands to the open ocean. Figure courtesy of Kelsea Jewell
Microbes are the major actors in the synthesis and degradation of all sorts of important molecules in environments. Cyanobacteria and algae in the oceans are responsible for the majority of photosynthesis on earth. They are the ultimate food source for most ocean creatures (including whales) and replenish the world's oxygen supply. Cyanobacteria also use carbon dioxide to synthesize their biological molecules and thus remove it from the atmosphere. Since carbon dioxide is a major greenhouse gas, its removal by cyanobacteria affects the global carbon dioxide balance and is an important mitigating factor in global warming.
In all habitats, microorganisms make nutrients available for the future growth of other living things by degrading dead organisms. Microbes are also essential in treating the large volume of sewage and wastewater produced by metropolitan areas, recycling it into clean water before discharging it into the environment. Less helpfully (from the view of most humans), termites contain microorganisms in their guts that assist in the digestion of wood, allowing the termites to extract nutrients from what would otherwise be indigestible. Understanding these systems helps us to manage them responsibly, and as we learn more, we will become ever more effective stewards.
Energy is essential for our industrial society, and microbes are important players in its production. A significant portion of natural gas comes from the past action of methanogens (methane-producing bacteria). Numerous bacteria are also capable of rapidly degrading oil in the presence of air. Special precautions during the drilling, transport, and storage of oil minimize the impact of these oil-degrading bacteria. In the future, microbes will find utility in the direct production of energy. For example, many landfills and sewage treatment plants capture the methane produced by methanogens to power turbines that produce electricity. Excess grain, crop waste, and animal waste can serve as nutrients for microbes that ferment this biomass into methanol or ethanol. These biofuels are presently added to gasoline and thus decreasing pollution. As part of the new energy initiative, scientists are engineering cyanobacteria to synthesize and excrete compounds used as biofuel. These microbes only need sunlight and seawater to grow, and therefore mass production of this biofuel should be inexpensive. Other research is investigating the possibility of using another photosynthetic bacterium, Rhodobacter sphaeroides, as a source of hydrogen production. One of these research efforts could one day power our cars, causing little to no pollution and having no net contribution to greenhouse gases.
Finally, we are increasingly taking advantage of the versatile appetite of bacteria to clean up environments that we have contaminated with crude oil, polychlorinated biphenyls (PCBs), and many other industrial wastes. This process is termed bioremediation and is a cheap and increasingly effective way of cleaning up pollution. Figure 1.7 shows before and after photographs of the clean-up of the Exxon Valdez oil spill in Prince William Sound. These techniques may become useful in the cleanup of the Deep Water Horizon oil spill in the Gulf of Mexico. Current research has shown that indigenous microbes are actively degrading the deep-sea oil plumes of that spill and may eventually detoxify much of that oil spill.

Figure 1.7. The Exxon Valdez oil spill. Microorganisms played an important role in removing many of the pollutants released during the Exxon Valdez oil spill in Prince William Sound. Interestingly, microbes were not added to the site, but the clean-up relied on bacteria from that environment. A nutrient solution was sprayed onto the oil to encourage the growth of oil-degrading microbes. Though this was one of the more successful methods used to clean up the oil, but no treatment removed all of the pollutants. (Source: http://www.battelle.org/environment/exxon-valdez.stm.)
Studying microbes helps us to understand the world around us
Microorganisms used in research have many useful properties. They grow on simple, cheap medium and often give rise to large populations in 24 hours. It is easy to isolate their genomic material, manipulate it in the test tube and then place it back into the microbe. Due to their large populations, it is possible to identify rare events and then, using powerful selective techniques, isolate interesting bacterial cells and study them. These advantages have made it possible to test hypotheses rapidly. Using microbes, scientists have expanded our knowledge about life. Below are a few examples.
Microorganisms have been indispensable instruments for unlocking the secrets of life. Our understanding of the molecular basis of heredity in organisms and its expression as proteins came through work on microorganisms. For an in-depth discussion on the molecular basis of heredity see the chapter on the Central Dogma. Since life at the molecular level is surprisingly similar, this understanding has helped us to learn about all organisms, including ourselves.
Some prokaryotes can grow under unimaginably harsh conditions that define the extreme limits of where life can exist. Some species can grow at near 100 °C in hot springs and even 113°C near deep-sea ocean vents. Figure 1.8 depicts such a deep-sea ocean vent. Others make their living at near 0 °C in freshwater lakes buried under Antarctica's ice. The ability of microbes to live under such extreme conditions is forcing scientists to rethink the requirements necessary to support life. Many now wonder if Jupiter's moon Europa may harbor living communities in waters deep below its icy crust. What may the rest of the universe hold?
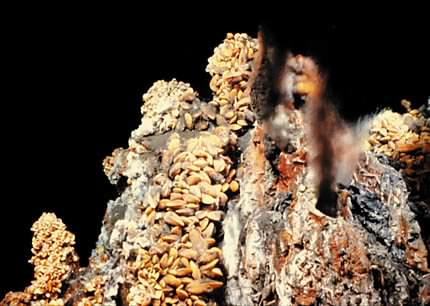
Figure 1.8. A deep sea ocean vent. Ocean vents are common in areas of the sea floor that have volcanic activity. Water seeps into cracks on the floor and encounters magma. The water absorbs inorganic nutrients from the magma and is heated. The superheated water then flows out of the magma, sometimes quite forcefully, back into the ocean. This hot water contacts the cold ocean water, causing it to cool and release many of its inorganic contents. This cloud of inorganic compounds is highly reduced and can serve as a source of energy for microorganisms. These microbes serve as a primary producer for an entire food chain. The picture shows one type of ocean vent called a black smoker. Water coming out of the vent can be >300°C. Figure courtesy of Woods Hole Oceanic Institute.
Until recently, while we could study specific types of bacteria, we lacked a cohesive classification system so that we could not readily predict the properties of one species based on the known properties of others. Visual appearance, which is the basis for the classification of large organisms, does not work with many microbes because there are few visible distinguishing characteristics for comparison between species even under the microscope. However, analysis of the genetic material of microbes in the past 20 years has allowed such classification. This reclassification spawned a revolution in our thinking about the evolution of bacteria and all other species. The emergence of a new system organizing life on earth into three domains is attributable to this pioneering work with microorganisms.
The fruits of basic research on microbes have been used by scientists to understand microbial activity and shape our modern world. Human proteins, especially hormones like insulin and human growth factor, are now produced in bacteria using genetic engineering. Our understanding of the immune system relied on using microbes as tools. Microorganisms also play a role in treating disease and keeping people healthy. Many of the drugs available to treat infectious disease originate from bacteria and fungi.
Lastly, microbes have informed us about our world through the tools they provide for molecular biology. Enzymes purified from bacterial strains are useful as tools to perform many types of analyses. With the rapid advance in DNA sequencing technology, it is now possible to determine the entire genetic sequence of any microbe desired in just a few weeks. With this information in hand, scientists can predict the capabilities of the microbe and manipulate DNA in useful ways. We now know the entire sequence of the human genome, except regions of repetitive DNA, which will hopefully lead to medical practices and treatments that improve health. We also know the entire genome sequences of hundreds of microbes, including those of many important pathogens. Analysis of this data will eventually lead to an understanding of the function of critical enzymes in these microbes and the development of tailor-made drugs to stop them. The rapid sequencing of SARS-CoV-2 allowed the development of vaccines and the tracking of various variants of the virus. In addition, scientists are mining the genomes of other microbes for novel antibiotics and combining various genes and pathways to create even more new drugs. Biotechnology firms can use this technology to create living synthetic factories that create many other useful compounds. The tools of molecular biology will also affect agriculture. For example, we now know the complete genome sequence of the plant Arabidopsis(a close relative of broccoli and cauliflower). Knowing plant sequences creates a better understanding of all plants and hopefully improvements in important crops.
Microbes have a profound impact on every facet of human life and everything around us. Pathogens harm us, yet other microbes protect us. Some microbes are pivotal in growing food crops, but others can kill the plants or spoil the produce. Bacteria and fungi eliminate the wastes produced in the environment, but also degrade things we would rather preserve. Clearly, they affect many things we find important as humans. In the remainder of this chapter, we take a look at how scientists came to be interested in microbes and follow a few crucial developments in the history of microbiology.
Key Takeaways
- Microbes make up the major portion of the biomass present on the earth. Therefore, the nutrients they eat and the products they form greatly influence the environment.
- Cyanobacteria and algae in the oceans are responsible for most photosynthesis and are a major sink for carbon dioxide, a greenhouse gas.
- Microbes release nutrients from dead organisms, making them available to the rest of the ecosystem. Some microbes play a role in the production of energy, while other microbes interfere with energy production.
- Microbes are useful tools in research because of their rapid life cycle, simple growth requirements, and small size. Due to this simplicity, microbes have been essential in understanding core questions in biology.
- Attempts to classify microorganisms have lead to a classification system that divides all organisms into three domains of life: Archaea, Bacteria, and Eukarya. Microbes provide tools for use in molecular biology. These tools have allowed scientists to make rapid progress in investigating many types of microorganisms.
1 - 5 Microscopes and Microbes
Learning Objectives
After reading this section, students will be able to...
- Describe how science is interdependent, with research building on earlier work.
- Describe the discovery of microorganisms.
- Explain the early classification work of Ferdinand Cohn.
- Describe the fundamental methods of Robert Koch's for cultivating microbes.
Before we begin the adventure that we call learning microbiology (it is an adventure! Really!), a look at the history of microbiology will help you understand the contributions of those who have come before. This perspective will hopefully give you an appreciation of their efforts and put the body of knowledge we will examine in the context of history. Keep in mind that microbiology is a relatively young science. It was only about 140 years ago that it became possible to seriously study microorganisms in the laboratory, with most of our understanding of microbes coming in the last 60 years.
The history of microbiology, like all human history, is not a catalog of linear progress but is more of an interweaving of the careers of bright individuals and their insights. Each discovery relied on previous ones and, in turn, spawned further inquiry. A web of interdependent concepts evolved through the work of scientists in many related disciplines and nations. Often the research of one individual impacted the efforts of another studying a completely different problem. Keep this in mind as you look at this history.
Below, we present several journeys through this web, mentioning some individuals who were particularly important in the progression. This history reflects our view of important events of the past but is by no means comprehensive. We will first look at the development of the techniques for handling microorganisms since everything else in microbiology depends upon these procedures. Next, we will examine how these techniques helped to settle an old debate, the question of spontaneous generation. Then, we will look at the history of infectious diseases. The science of microbiology had its most significant early impact on human health, uncovering the cause of the major killers of the day and then methods to treat them. As microbiology matured, scientists began to look at what non-pathogenic microbes were doing in the environment, and we will look a bit at the history of general microbiology. Finally, the chapter will examine the events that lead to the understanding of life at the molecular level and the profound impact this has had on microbiology and society in general.
Early observations
For years, the existence of microorganisms was suspected but could not be proven since bacteria were too small to be seen with the naked eye. It took the microscope to expose their tiny world, and that instrument has been linked to microbiology ever since. In 1664, Robert Hooke devised a compound microscope and used it to observe fleas, sponges, bird feathers, plants, and molds, among other items. His work was published in Micrographia and became a popular and widely read book at the time.
Several years later, Anton van Leeuwenhoek, a fabric merchant and amateur scientist (or "natural philosopher" as such people called themselves), became very adept at grinding glass lenses to make telescopes and microscopes. While crude by modern standards, his were a technical marvel for the time, able to magnify samples greater than 200-fold - a reproduction of a Leeuwenhoek microscope is shown in Figure 1.9. They also produced clearer images than the compound microscopes of the time. By peering through his microscope, Leeuwenhoek observed tiny organisms or "wee animacules" as he called them. He spent months looking at every kind of sample he could find and eventually submitted his observations in a letter to the Royal Society of London, causing a sensation. Hooke confirmed the findings of Leeuwenhoek, and his affirmative assessment garnered wide acceptance. Leeuwenhoek received many honors for his work and lived well into his 9th decade.
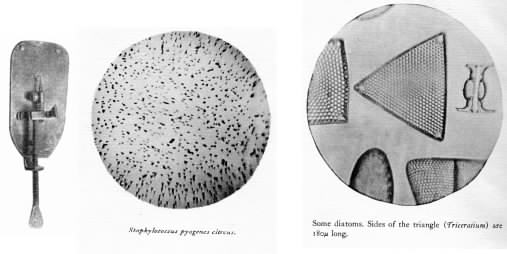
Figure 1.9. The Van Leeuwenhoek microscope.. The left panel shows a replica of a Leeuwenhoek microscope . The photomicrographs in the center and right were taken in the early 20th century through one of Leeuwenhoek's microscope. (Source: The Leeuwenhoek Letter. Society of American Bacteriologists. Baltimore. 1937.)
Surprisingly, there was no follow-up work on these findings for over 200 years. Human societies had neither the technical prowess nor the inclination to develop the science of microorganisms. It was not until the rise of the industrial revolution that governments and people dedicated the financial and physical resources to understand these small inhabitants of our world.
With the development of better microscopes in the 19th century, scientists returned to an examination of microorganisms. After finishing his education, Ferdinand Julius Cohn convinced his father to lay down the large sum necessary to purchase a microscope for him, one better than that available at the university in Breslau, then part of Germany. He used it to sift through the world of the microbe and made many observations of eukaryotic microorganisms and bacteria. Cohn published his landmark papers on the cycling of elements in nature in Ueber Bakterien in 1872. Other work, including a microbial classification scheme with descriptions of Bacillus, appeared in the first volume of a journal he founded, Beitraege Zur Biologie der Pflanzen.. Cohn's work with microscopes popularized their use in microbiology. His work inspired many other scientists to examine microbes. Cohn's encouragement of Robert Koch, a German physician by training, began the field of medical microbiology.
Robert Koch developed many microbiological techniques
Robert Koch, pictured in Figure 1.10, searched for the causes of many diseases. Through these investigations, he and his laboratory developed many classic microbial techniques. He used adaptations of the staining methods of Carl Weigert to begin the process of distinguishing microbes and identifying pathogens. His lab was the first to isolate a disease-causing organism.
Figure 1.10. Robert Koch. The German Physician Robert Koch. Drawing by Tammi Henke
A major contribution to bacterial techniques was the development of methods using solid medium. For example rich medium or minimal medium for the cultivation of bacteria. Koch suspected that microbes caused some diseases. However, to test this idea, he needed to isolate the causative agent. Almost all samples from diseased animals or any natural surface contained many different microbes, and it was impossible to tell which one was the problem. Koch needed a method to separate these different bacteria. The most common isolation method was to continually dilute a sample in liquid broth, hoping that only one microbe would eventually be present at high enough dilution. A problem with this method is that the scientist would isolate only the most populous microbe, but that might not be one causing the disease. There were other technical problems with such a liquid-based system, so a solid medium would seem to provide distinct advantages. Koch had tried gelatin for these experiments with unsatisfactory results. Building on the work of Brefeld and Schroeter, Koch used potato slices as a solid medium and observed that a boiled potato left in the open air would develop tiny circular raised spots.
Examination of these spots revealed they were piles of microorganisms, and each spot had just one type of microbe in it. He realized that these colonies were pure bacterial cultures and probably arose from a single microbe species from the air that landed on the potato. By boiling a potato, slicing it with a hot knife, and keeping it in a sterile container with a lid, Koch could keep the potato sterile. But if he smeared a sample from a diseased animal across the potato, colonies arose, each being pure isolates. By then testing these isolates in animals, Koch was able to isolate the cause of anthrax, Bacillus anthracis.
Potatoes failed to support the growth of many microorganisms, and Koch and his laboratory were constantly frustrated by the lack of a suitable solid medium. Walter Hesse joined Koch's laboratory to study air quality, showing remarkable attention to detail and patience in his work. His wife, Angelina Fannie Hesse, along with raising their three sons, also would assist her husband with his research in the laboratory. Walter was attempting to do his air quality experiments using a medium containing gelatin as the solidifying agent. In the summertime, temperatures would often rise above the melting point of gelatin. In addition, microbes would often grow in the cultures that were capable of degrading gelatin, and in both cases, this would cause liquefaction of the medium, ruining the experiments. One day while eating lunch, the frustrated scientist asked Lina (as he called her) why her jellies and puddings stayed solid even in the hot summer temperatures. She told him about agar-agar, a heat resistant gelling agent that she had learned about while growing up in New York from a Dutch neighbor who had emigrated from Java.
Development of the new agent by Angelina and Walter led to a resounding success. Few microbes can degrade agar, and it melts at 100 °C yet remains molten at temperatures above 45 °C. This property allows the mixing of the agar with heat-sensitive nutrients and microbes. After solidification, it does not melt until a temperature of 100 °C is again attained, facilitating the easy cultivation of pathogens. It can also be stored for long periods, allowing the cultivation of slow-growing microorganisms. Any broth can be mixed with agar, giving great flexibility in the kinds of media that an investigator can make. Thus, researchers could suddenly cultivate many more types of microbes. Agar is still used today as a solidifying agent.
Koch's laboratory also developed methods of pure culture maintenance and aseptic technique. Aseptic technique involves the manipulation of pure cultures in a manner that prevents contamination by outside microorganisms. Equally important, aseptic technique prevents their spread into the environment. Remember that Koch was studying some of the most devastating microbial pathogens of the period, and their release could potentially cause disease in the scientists working on them. These procedures were also absolutely critical because they allowed careful study of pure microorganisms, making it possible to identify the role of each microbe in a given situation.
Another problem in cultivating microbes was solved by Julius Petri while working in Koch's laboratory. Solid medium was poured on glass plates and allowed to spread and harden. Once cooled it allowed a solid surface for streaking. However, creating these plates required great care since exposure to the air (and the microbes in it) often leads to contamination. In addition, to prevent contamination of plates during incubation, researchers used a cumbersome bell jar. If one wanted to view samples, the plate had to be removed from the jar, further exposing it to unwanted microbes in the air. In 1887 Petri developed shallow glass dishes, with one having a slightly larger diameter than the other. Medium is poured into the smaller dish, and the larger one serves as a cover. This simple device solved all of the above problems and took on the inventors name, the Petri plate.
These same techniques are essential in studying all microorganisms. Collectively the above techniques have been used to isolate and identify thousands of different microorganisms. As a testament to the significance of their achievement, these techniques are practiced with remarkably little change in every laboratory that works with microorganisms today. Table 2 lists some of the early advances that helped to develop the practice of microbiology.
Table 2. The development of early techniques in microbiology.
| Year | Event |
| 1664 | Robert Hooke is the first to use a microscope to describe the fruiting structures of molds. He also coined the term cell when using a microscope to look at cork, as the dead plant material in cork reminded him of a jail cell. |
| 1673 | Anton van Leeuwenhoek, a Dutch tradesman and skilled lens maker, is the first to describe microbes in detail. |
| 1872 | Ferdinand Julius Cohn publishes landmark paper on bacteria and the cycling of elements. In it is an early classification scheme that uses the name Bacillus. |
| 1872 | Oscar Brefeld reports the growth of fungal colonies from single spores on gelatin and the German botanist Joseph Schroeter grows pigmented bacterial colonies on slices of potato. |
| 1877 | Robert Koch develops methods for staining bacteria, photographing, and preparing permanent visual records on slides. |
| 1881 | Koch develops solid culture media and the methods for obtaining pure cultures of bacteria. |
| 1882 | Angelina Fannie and Walther Hesse in Koch's laboratory develop the use of agar as a support medium for solid culture. |
| 1884 | Hans Christian Gram develops a dye system for identifying bacteria [the Gram stain]. |
| 1887 | First report of the petri plate by Julius R. Petri. |
| 1915 | M. H. McCrady establishes a quantitative approach for analyzing water samples using the most probable number, multiple-tube fermentation test. |
Key Takeaways
- Science is interdependent and new discoveries depend upon earlier contributions from many other scientists.
- Microbes were first seriously described in the 17th century by Robert Hooke and Anton van Leeuwenhoek using simple microscopes.
- Ferdinand Cohn continued this work many years later, making a first systematic attempt at classifying them.
- Robert Koch developed techniques for handling microbes and many of these methods for cultivating microbes are still used today.
1 - 6 Spontaneous generation was an attractive theory to many people, but was ultimately disproven.
Learning Objectives
After reading this section, students will be able to...
- Explain why people believed in the concept of spontaneous generation, the creation of life from organic matter.
- Describe the experiment by Francesco Redi disproved spontaneous generation that disproved spontaneous generation for macroorganisms.
- Explain how did John Needham's experiment re-ignited the debate about spontaneous generation for microorganisms.
- Describe the swan-neck flask experiment of Louis Pasteur and why this ended the debate about spontaneous generation.
Spontaneous generation hypothesizes that some vital force contained in or given to organic matter can create living organisms from inanimate objects. Spontaneous generation was a widely held belief throughout the middle ages and into the latter half of the 19th century. Some people still believe in it today. The idea was attractive because it meshed nicely with the prevailing religious views of how God created the universe. There was a strong bias to legitimize the idea because this vital force was considered a strong proof of God's presence in the world. Proponents offered many recipes and experiments in proof. To create mice, mix dirty underwear and wheat grain in a bucket and leave it open outside. In 21 days or less, you would have mice. The real cause may seem obvious from a modern perspective, but to the supporters of this idea, the mice spontaneously arose from the wheat kernels.
Another often-used example was the generation of maggots from meat left in the open. Francesco Redi revealed the failing here in 1668 with a classic experiment. Redi suspected that flies landing on the meat laid eggs that eventually grew into maggots. To test this idea, he devised the experiment shown in Figure 1.11. Here he used three pieces of meat. Redi placed one piece of meat under a piece of paper. The flies could not lay eggs onto the meat, and no maggots developed. The second piece was left in the open air, resulting in maggots. In the final test, Redi overlayed the third piece of meat with cheesecloth. The flies could lay the eggs into the cheesecloth, and when he removed this, no maggots developed. However, if Redi placed the cheesecloth containing the eggs on a fresh piece of meat, maggots developed, showing it was the eggs that "caused" maggots and not spontaneous generation. Redi ended the debate about spontaneous generation for large organisms. However, spontaneous generation was so seductive a concept that even Redi believed it was possible in other circumstances.
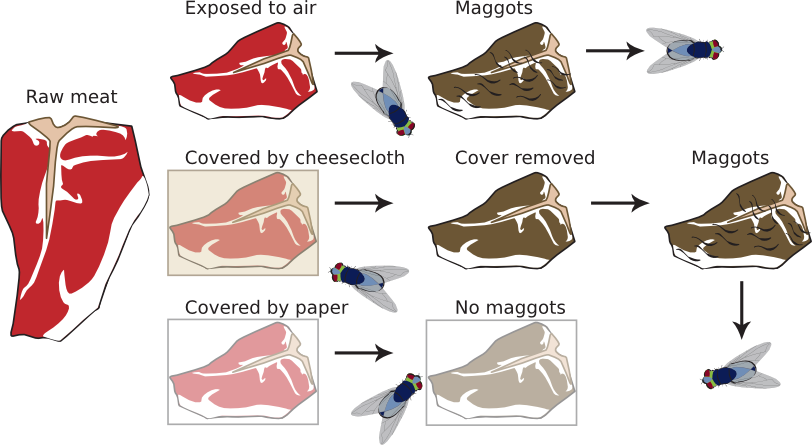
Figure 1.11. The Redi experiment.. Using several pieces of meat, paper and cheesecloth, Francesco Redi produced compelling evidence against the theory of spontaneous generation. One of the strong points of this experiment was its simplicity, which allowed others to easily reproduce it for themselves. See the text for details of the experiment.
The concept and the debate were revived in 1745 by the experiments of John Needham. It was known at the time that heat was lethal to living organisms. Needham theorized that if he took chicken broth and heated it, all living things in it would die. After heating some broth, he let a flask cool and sit at a constant temperature. The development of a thick turbid solution of microorganisms in the flask was strong proof to Needham of the existence of spontaneous generation. Lazzaro Spallanzani later repeated the experiments of Needham, but removed air from the flask, suspecting that the air was providing a source of contamination. No growth occurred in Spallanzani's flasks, and he took this as evidence that Needham was wrong. Proponents of spontaneous generation discounted the experiment by asserting that the vital force needed air to work properly.
It was not until almost 100 years later that the great French chemist Louis Pasteur, pictured in Figure 1.12, put the debate to rest. He first showed that the air is full of microorganisms by passing air through gun cotton filters. The filter trapped tiny particles floating in the air. By dissolving the cotton with an ether/alcohol mixture, the particles were released and then settled to the bottom of the liquid. Inspection of this material revealed numerous microbes that resembled the types of bacteria often found in putrefying media. Pasteur realized that if these bacteria were present in the air, they would likely land on and contaminate any exposed material.

Figure 1.12. Louis Pasteur. The French microbiologist Louis Pasteur. Drawing by Tammi Henke
Pasteur then entered a contest sponsored by The French Academy of Sciences to disprove the theory of spontaneous generation. Similar to Spallanzani's experiments, Pasteur's experiment, pictured in Figure 1.13, used heat to kill the microbes but left the end of the flask open to the air. In a simple but brilliant modification, he heated the neck of the flask to melting and drew it out into a long S-shaped curve, preventing the dust particles and their load of microbes from ever reaching the flask. After prolonged incubation, the flasks remained free of life and ended the debate for most scientists.
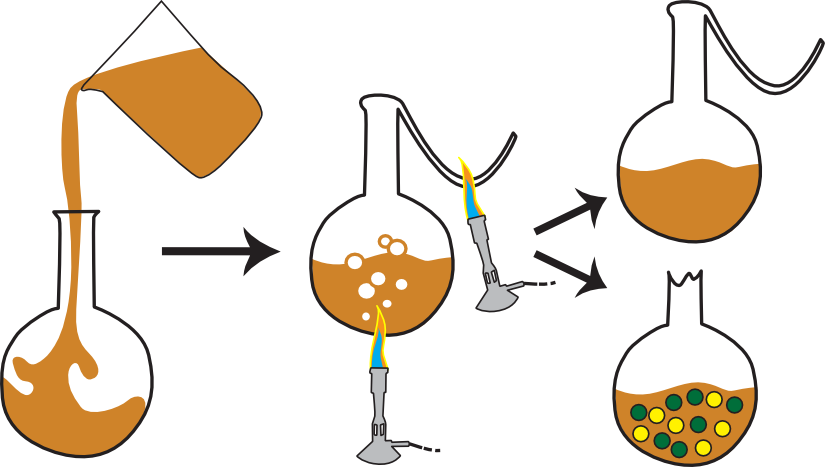
Figure 1.13. The swan neck flask experiment. Pasteur filled a flask with medium, heated it to kill all life, and then drew out the neck of the flask into a long S shape. This prevented microorganisms in the air from easily entering the flask, yet allowed some air interchange. If the swan neck was broken, microbes readily entered the flask and grew
A final footnote on the topic was added when John Tyndall showed the existence of heat-resistant spores in many materials. Boiling does not kill these spores, and their presence in chicken broth, as well as many other materials, explains the results of Needham's experiments.
While this debate may seem silly from a modern perspective, remember that the scientists of the time had little knowledge of microorganisms. Koch would not isolate microbes until 1881. The proponents of spontaneous generation were neither sloppy experimenters nor stupid. They did careful experiments and interpreted them with their own biases. Detractors of the theory of spontaneous generation were just as guilty of bias but in the opposite direction. It is somewhat surprising that Pasteur and Spallanzoni did not get growth in their cultures since the sterilization conditions they used would often not kill endospores. Luck certainly played a role. It is important to keep in mind that the discipline of science is performed by humans with all the fallibility and bias inherent in the species. Only the self-correcting nature of the practice reduces the impact of these biases on generally held theories. Spontaneous generation was a severe test of scientific experimentation because it was such a seductive and widely held belief. Yet, even spontaneous generation was overthrown when the weight of careful experimentation argued against it. Table 1.3 lists important events in the spontaneous generation debate.
Table 1.3 Events in spontaneous generation
| Year | Event |
| 1668 | Francesco Redi attacks spontaneous generation and disproves it for large organisms |
| 1745 | John Needham adds chick broth to a flask and boils it, lets it cool and waits. Microbes grow and he proposes it as an example of spontaneous generation. |
| 1768 | Lazzaro Spallanzani repeats Needham's experiment, but removes all the air from the flask. No growth occurs. |
| 1859 | Louis Pasteur's swan-neck flasks show that spontaneous generation does not occur. |
| 1870 |
Thomas H. Huxley gives his "Biogenesis and Abiogenesis" lecture. The speech offered powerful support for Pasteur's claim to have experimentally disproved spontaneous generation. |
| 1877 | John Tyndall publishes his method for fractional sterilization, showing the existence of heat-resistant bacterial spores. |
Key Takeaways
- For many centuries many people believed in the concept of spontaneous generation, the creation of life from organic matter.
- Francesco Redi disproved spontaneous generation for large organisms by showing that maggots arose from meat only when flies laid eggs in the meat.
- Spontaneous generation for small organisms again gained favor when John Needham showed that if a broth was boiled (presumed to kill all life) and then allowed to sit in the open air, it became cloudy.
- Louis Pasteur ended the debate with his famous swan-neck flask experiment, which allowed air to contact the broth. Microbes present in the dust were not able to navigate the tortuous bends in the neck of the flask.
1 - 7 Microbes are discovered to cause disease
Learning Objectives
After reading this section, students will be able to...
- Describe child-bed fever and how Ignaz Semmelweis showed physicians spread it and changing their behavior could prevent it.
- Explain the causes of sour wine and how Louis Pasteur associated this with infectious disease.
- Describe the isolation of the first disease microbe by how Robert Koch and the methods he used to achieve this.
- Describe the discovery of viruses and the scientists involved
Many philosophers and physicians long suspected that living things were the agents of disease. In volume 6 of his epic poem De Rerum Natural (On the Nature of the Universe) written sometime around 50 B.C., Titus Lucretius Carus speculates about invisible atoms causing disease. This poem was only one idea among many. Some thought that an imbalance in humors caused illness, while others felt that supernatural forces were at work. The prevailing theory held by most doctors of the 19thcentury was that chemical toxins were carried from an ill patient to others, causing them to contract the same malady. The bacteria that were present were seen as a symptom of the disease and not its cause.
Ignaz Semmelweis, pictured in Figure 1.14, a Hungarian physician working in Vienna, made the first breakthrough in the true nature of disease. He realized that asepsis in obstetrical wards could prevent the transmission of childbirth fever from patient to patient. He, therefore, instigated a policy for all attending physicians to wash their hands with chloride of lime (a mixture of calcium chloride hypochlorite, CaCl(OCl); calcium hypochlorite, Ca(OCl)2; and calcium chloride, CaCl2) between patients. This innovation dropped the mortality rate from 18% to 2.4%.

Figure 1.14. Ignaz Semmelweis. The Hungarian Ignaz Semmelweis. Drawing by Tammi Henke
Ignaz became a vigorous proponent of his ideas, but the Hungarian doctor's efforts were opposed by many who could not accept that physicians themselves could be responsible for spreading bacterial infection. Ridicule of his idea caused him to move from Vienna to Pest, Hungary and ultimately played a role in a nervous breakdown. Ironically, Semmelweis died from an infection that he contracted during surgery while recovering from his nervous breakdown.
Before his death, he published his ideas in a paper The Cause, Concept, and Prophylaxis of Childbed Fever in 1861. The paper's poor writing contributed to the obscurity of the ideas, with physicians ignoring them for 17 years. The delay raises an interesting point about the culture of science. Radical ideas, even those that are correct and can save lives, are sometimes ignored. It takes time to overcome the dogma of the day. The personalities involved and the negative light it might throw on past practices play a large role in the rate of acceptance of a new idea.
In a seemingly unrelated event, Louis Pasteur found something interesting while working on wine souring, a problem where wine fermentations produce a sour taste and very little alcohol. Fermentation of alcoholic beverages was thought to be a simple chemical reaction. Heating, common in most beverage preparations, was thought to cause the breakdown of sugar into alcohol. The wine industry in France asked for Pasteur's help because wine souring was pushing it close to ruin. His work on wine-making revealed that various yeast strains actually perform the process of converting sugar to alcohol. He then showed that the wine was going bad because a contaminating microbe generated lactic acid instead of alcohol from the sugar. This controversial idea gained credibility when Pasteur solved the problem by heating the wine and killing the contaminant. The heating process was named pasteurization in his honor and is still widely used today. In a brilliant step of generalization, Pasteur realized that souring of wine and infectious disease shared a common thread in that they both might involve infection by a microorganism. His suggestion that microbes cause disease became known as the germ theory of disease.
Joseph Lister became aware of Semmelweis's work and, together with Pasteur, realized the true nature of disease. He then recognized that he could use this idea to help his surgery patients. At this time, major injuries, broken bones, or surgery would often result in infection of the damaged area, sometimes leading to amputation or death. Lister found he could greatly reduce the number of microorganisms on wounds and incisions by using bandages treated with phenic acid, a compound that killed microorganisms (phenic acid, now known as phenol, is the active ingredient in Listerine). During surgery, he began the practice of spraying the wound with a fine mist of phenic acid to kill microbes. These practices greatly reduced the rate of infection and mortality of surgery patients, lending further credence to the germ theory of disease.
In 1876, Robert Koch provided definitive proof of the germ theory by isolating the cause of anthrax and showing it to be a bacterium. From this came the development of Koch's Postulates, a set of rules for assigning a microbe as the cause of a disease:
- The specific organism should be shown to be present in all cases of animals suffering from a specific disease but should not be found in healthy animals.
- The specific microorganism should be isolated from the diseased animal and grown in pure culture on artificial laboratory media.
- This freshly isolated microorganism, when inoculated into a healthy non-immune laboratory animal, should cause the same disease seen in the original animal.
- The microorganism should be reisolated in pure culture from the experimental infection.
The postulates are Koch's most famous contribution to science, and it is a testament to the utility of these postulates that they are stilled used today to discover the cause of new emerging diseases. Koch went on to apply these principles to study many other diseases, including tuberculosis, cholera, and sleeping sickness. Koch's postulates do have limitations and do not apply to all diseases. For example, if a disease-causing microbe has humans as its sole host and has a significant possibility of causing death, it would be unethical to apply this microbe to test humans as dictated by postulate. Also, it is not always possible to obtain a disease-causing microbe in pure culture.
While attacking the problem of disease, Koch developed the tools for obtaining pure cultures. Advances in science often come from innovations in the available technology. Robert Koch was an influential microbiologist because his pioneering work in the isolation and characterization of bacterial diseases helped identify the causes of many of the disorders plaguing humanity. Further work by other scientists then began the long road to conquering them.
Viruses are shown to parasitize organisms
The ensuing years brought numerous discoveries about the nature of disease. In the latter half of the 19thcentury, scientists discovered the causative agents for anthrax, tuberculosis, gonorrhea, diphtheria, and many more. Researchers made great strides in understanding disorders caused by bacteria, including the solidification of the germ theory of disease. However, some illnesses seemed not to have bacterial origins. Microscopic examination of sera from ill patients revealed no organisms, and the causative agents could not be grown on any known medium. Yet if these sera were injected into a susceptible host, disease resulted.
In 1886 John Brown Buist devised a method for staining and fixing liquid from a cowpox vesicle. Observations of this slide showed tiny bodies that he believed were spores. Although he did not realize it, he was the first person to see (and photograph) a virus. It was not until the middle of the twentieth century, with the invention of the electron microscope, that we understood the true shape and structure of viruses.
In 1884 Charles Chamberland in Pasteur's laboratory created an unglazed porcelain filter with pores much smaller than bacteria (0.1-1 µm). It was possible to pass a solution containing bacteria through these filters and completely remove them from the solution. This innovation enabled the creation of sterile medium without heat and became a standard test for removing all bacteria, especially when testing transmission of disease. By removing the microbes, Chamberland demonstrated that an infectious agent, and not the solution, was causing the illness. This concept seemed reasonable until scientists began to investigate a tobacco infection.
Tobacco mosaic disease is characterized by light and dark green areas on plant leaves in a mosaic pattern. The disease stunts the growth of the plant, therefore reducing yields. Europeans recognized the effects of this disease soon after farmers introduced tobacco from the New World in the 17th century. Adolph Mayer first described the transmission of the disease by injecting fluid from a diseased plant into a healthy one. Nine out of ten times, the healthy plant would become heavily diseased. In 1892 Dmitrii Ivanowski extended this research by the shocking discovery that the causative agent could pass through Chamberland's porcelain filter. He reported his findings, but the idea that the causative agent could pass through the filter was so troublesome that Ivanowsky attributed the phenomenon to a cracked filter or to small spores that passed through the pores. It was Martinus Beijerinck in 1898 who realized the true nature of these particles, making the intellectual leap that the causative agent of the disease must be so small as to pass through the filter known to trap all bacteria. He coined the term contagium vivum fluidum a contagious living fluid.
Some scientists thought these agents were toxins in the fluids of the hosts, but further study revealed that tobacco mosaic disease could cause illness through many transfers from plant to plant. A toxin might cause damage on the first plant, but subsequent transfers should make it so dilute that it would no longer have any effect. These filterable entities were different from bacteria and appeared to depend on their host to multiply. The term ultrafiltrable viruses and later just viruses (virus means poison in Latin) was coined to describe these tiny pathogens.
Bacteria are also vulnerable to viruses. Frederick Twort described bacterial in 1915 and independently by Felix d'Herelle in 1917. Felix d'Herelle was studying a plague of locusts in Mexico when he noticed that a microorganism was killing the locusts. During experiments to characterize this microbe, d'Herelle noticed clear, circular spots two or three millimeters in diameter on cultures growing on agar. At this point d'Herelle studied the spots enough to determine that they came from an agent small enough to pass through a porcelain filter. He dropped the investigation but recalled these observations while studying dysentery in 1915. The illness affected soldiers fighting in World War I, and he soon determined the cause to be the bacterium Shigella dysentery. In the process of investigating Shigella, similar areas of clearing were observed. d'Herelle realized that something in these areas of clearing was killing the bacterium. He eventually determined these were viruses of bacteria, coining the term bacteriophage (devourer of bacteria). Table 1.4 lists these and other significant events in the discovery of the cause of disease.
Table 1.4 Important events in the discovery of the cause of disease.
| Year | Event |
| 1840s | Ignaz Semmelweis shows that hand washing between visiting mothers can prevent childbirth fever. |
| 1854 | Dr. John Snow studies a cholera outbreak in the Soho neighborhood of London and determines it was caused by contaminated water at the Broad Street pump. His methods found the field of epidemiology. |
| 1857 | Louis Pasteur develops the germ theory. |
| 1867 | Joseph Lister develops the use of phenic acid (phenol) to treat wounds and for antiseptic surgery. |
| 1873 | Gerhard Henrik Armauer Hansen discovers the leprosy bacillus (Mycobacterium leprae) and demonstrates that leprosy is a contagious disease and not inherited as was the popular belief. In many countries leprosy is still called Hansen's disease in his honor. |
| 1876 | Robert Koch and Cohn identify a bacterium, Bacillus anthracis as the cause of anthrax and publish their research. |
| 1882 | Koch isolates the Tuberculosis bacillus, Mycobacterium tuberculosis. |
| 1884 | Koch puts forth his postulates, which are standards for proving that a microorganism is the cause of a disease. Application of Koch's postulates continues to reveal the association of many diseases with pathogens. |
| 1886 | John Brown Buist is the first person to see a virus. |
| 1892 | Dmitri Ivanowski publishes the first evidence of the filterability of a pathogenic agent, the virus of tobacco mosaic disease. |
| 1899 | Martinus Beijerinck recognizes the unique nature of Ivanowski's discovery. He coins the term contagium vivum fluidum - a contagious living fluid. |
| 1899 | Friederich Loeffler and Paul Frosch discover that foot and mouth disease is also caused by a filterable agent. |
| 1915-1917 | Frederick Twort and Felix d'Herelle discover bacterial viruses. |
| 1918 | In the fall of 1918, as World War I was ending, an influenza pandemic of unprecedented virulence swept the globe, leaving some 40 million dead in its wake. A search for the responsible agent began in earnest that year, leading to the first isolation of an influenza virus by 1930. |
| 1957 | D. Carleton Gajdusek proposes that a slow virus is responsible for the wasting disease kuru. In subsequent years several diseases are shown to be caused by slow viruses (later renamed prions) including mad cow disease. |
Key Takeaways
- Ignaz Semmelweis showed that child-bed fever was spread by physicians and could be prevented by careful hand washing with chloride of lime.
- Louis Pasteur, while working on sour wine, discovered that unwanted microbes were infecting the wine. He correctly deduced that similar infections with harmful microbes caused infectious disease.
- Robert Koch was the first to isolate a disease-causing microbe, Bacillus anthracis. In the process, he developed techniques and standard protocols for defining the cause of a disease.
- John Brown Buist was the first to observe a virus, the cowpox virus, although he did not realize it.
- Dmitrii Ivanowski discovered that the cause of tobacco mosaic disease could pass through a porcelain filter, but it was Martinus Beijerinck who correctly deduced that the particle causing the disease was too small to be a bacteria and would later be known as a virus.
- Frederick Twort and Felix d'Herelle each discovered bacteriophage, viruses that infect bacteria.
1 - 8 Treating Disease
Learning Objectives
After reading this section, students will be able to...
- Describe the practice of variolation.
- Describe Edward Jenner's role in developing the first vaccine.
- Explain how Pasteur discovered vaccination as a general practice.
- Describe what Paul Ehrlich did for 17 years.
- Explain the contribution of Alexander Fleming.
The power of vaccination is discovered
As soon as physicians realized that microbes cause illness, the search was on for ways to kill or to prevent them from causing disease. In this section and the next, we will look at two series of events that illustrate the emergence of modern treatment of infectious disease: the development of vaccines and the discovery of antimicrobial compounds.
Smallpox was a feared disease throughout human history and justifiably so. It was highly contagious, and almost everyone eventually became infected. Mortality rates were as high as 25% in adults and closer to 40% in children. Those who did survive often had scarring due to the blister-like pustules that form on the skin, but they obtained life-long immunity to the disease.
As far back at the 11thcentury in India and China, doctors realized that liquid from the pustules of a smallpox victim, when scratched on the skin of a healthy patient, would most often cause mild disease. This intentional infection, termed variolation, would also give life-long protection against the virus. Lady Mary Wortley Montague, the ambassador's wife to the Ottoman Empire, introduced variolation to England in 1721. It quickly became a popular practice throughout Europe. Washington even began variolating the Continental Army in 1776.
Variolation had some deleterious side effects. Serious skin lesions inevitably resulted at the site of inoculation, often accompanied by a generalized rash or even a full case of smallpox. The fatality rate from variolation was 1 to 2 %. Today we would find this fatality level to be unacceptable, but at the time, this risk still represented a significant advance.
In 1796, Edward Jenner, an English country physician pictured in Figure 1.15, went in search of a more predictable and safer method of protection against the disease. He noticed that milkmaids rarely contracted smallpox. Further investigation revealed they often contracted cowpox from their charges. Jenner hypothesized that cowpox was related to smallpox, and contraction of the former would protect against the latter. In a classic experiment (and one that would land you in jail today), Jenner inoculated a young patient with cowpox and later challenged him with smallpox. The boy did not become ill, and Jenner was responsible for creating a safer method of protection against smallpox. It is important to stress that the nature of these diseases and their viruses would not be known for over 100 years. Jenner was ahead of his time.

Figure 1.15. Edward Jenner. The English physician who developed a safer method for protecting against smallpox. We would now call this a vaccine. Drawing by Tammi Henke.
Beginning around 1876 Pasteur's studies on chicken cholera led to the development of vaccines to fight the disease. Cholera was a serious problem since it could spread through a barnyard and wipe out a flock in as little as three days. Contaminated food or animal excrement spread the disease. Pasteur identified the cholera bacillus and grew it in pure culture. When injected with it, a chicken invariably died within 48 hours.
Then, as often happens in scientific research, luck intervened. During the heat of the summer, Pasteur returned to Paris and left the cholera cultures used for infection stored on the shelves of his laboratory in Arbois, France. Upon returning, something had happened to the cultures; they no longer caused disease when tested in chickens. With some impatience for the time they were wasting, his group set to work making new cultures of the bacillus and tested these batches on both new birds and those previously inoculated with the ineffective strain. To their amazement, the previously injected birds were unaffected by the fresh bacillus culture, while the new birds all died. Pasteur immediately realized that this was similar to the studies of Jenner.
Pasteur then developed a method for creating cultures that would confer immunity, but not cause disease. Sometimes this involved growing the microbe in a medium in the laboratory where they would spontaneously lose their virulence. In other cases, it involved multiple passes through a susceptible host. For example, Pasteur attenuated the rabies virus by passing it through rabbits. In honor of Jenner's accomplishments Pasteur coined the term vaccination (vacca = cow in Latin) for the process of immunization against disease. In this and several of Pasteur's other discoveries, luck played a part, but it was only helpful because he tenaciously pursued odd results and had the insight to arrive at important conclusions. In Pasteur's famous words, In the field of observation, chance favors only the prepared mind
Pasteur's technique of weakening a strain by a damaging treatment or passing it through a susceptible host was termed attenuation and resulted in the creation of vaccines against anthrax, plague, yellow fever, rabies, and many other diseases. Scientists have developed many vaccines over the years and children today receive various shots, significantly decreasing infant mortality and illness.
Antimicrobial compounds are developed to kill microorganisms
We now pick up another thread through the web of microbial history that began with the work of Paul Ehrlich. By 1885, it was becoming clear that the causative agents of many illnesses were microorganisms. As scientists manipulated these microbes in the lab, they found that certain dyes and other compounds could inhibit their growth. Paul Ehrlich began the search for chemicals that kill the microbe, but not the patient, thus curing the illness.
One of the diseases Ehrlich hoped to cure was syphilis, which had reached epidemic proportions in Europe. Little did he realize it would be a 17-year odyssey before developing salvarsan, the first effective chemotherapeutic agent. Salvarsan was the 606thchemical he tried. This arsenic compound effectively kills Treponema pallidum, the causative agent of syphilis.
However, the treatment had many problems, causing long-lasting health complications for those who used it. In addition, despite Treponema being quite sensitive to salvarsan, the physician had to administer it intravenously for optimum effectiveness. Intravenous injection was a recent development, and many doctors were leery of trying the procedure. In London, a young physician by the name of Alexander Fleming, then in the Army Medical Corps, was one of the few that was willing to treat patients. Fleming even got the nickname private 606 from his burgeoning practice. His work validated the effectiveness of salvarsan against syphilis and convinced others to administer the treatment.
Fleming, pictured in Figure 1.16, was a physician by training but spent most of his time studying bacteria. His success with salvarsan motivated him to search for other antibacterial agents. His first discovery was lysozyme, an enzyme produced by many organisms, including humans, which lysed many bacteria. This enzyme is not useful as a therapeutic agent because it is difficult to administer as a drug. Still, Fleming did develop titration methods and assays that would become very useful in his future work.

Figure 1.16. Alexander Fleming. The young English physician who discovered the antibiotic penicillin. Drawing by Tammi Henke.
Fleming's arguably most important contribution to science is his discovery of penicillin. In September of 1928, before leaving on a summer holiday, Fleming streaked some plates of Staphylococcus aureus and left them to incubate until his return. In an improbable set of circumstances, the beginning of the holiday was cold, allowing some contaminating mold spores (that had blown in from a nearby window) to grow up on some of the plates. The temperature then increased encouraging the growth of the Staphylococcus. When confronted with a contaminated plate, many experimenters look for the trash bin, but Fleming instead spent some time examining it. The fungus had a zone of clearing around it where the Staphylococcus colonies would not grow, suggesting the fungus was producing an antibacterial compound that had diffused into the medium. Intrigued, he cultured the fungus, a Penicillium mold, and eventually isolated a soluble extract that could kill bacteria and treat a localized infection. He called the new compound penicillin after the mold from which it came. However, due to available technology, it was challenging to prepare a solution that could be used throughout the body without causing problems.
World War II added greater urgency to the search for compounds that could fight infectious disease. If they survived the initial injury, wounded soldiers would often develop life-threatening infections, and there were no effective drugs to combat them. In 1939 Howard Florey and Ernst Chain began a systematic study of antimicrobial compounds to develop treatments for these soldiers and ran across Fleming's report written nine years earlier. They were now able to purify the compound completely and describe its high potency against microbes. The availability of penicillin during World War II saved countless lives. The rediscovery of penicillin touched off a search for other microbes producing substances that could kill or inhibit microbes, leading to the discovery of many more antimicrobials. With Florey and Chain, Fleming was awarded the Noble Prize in Medicine and Physiology in 1945.
In the ensuing decades, the search and discovery of numerous antimicrobial compounds, combined with the development of vaccines, has eliminated many of the deadly diseases that plagued humankind. While we now realize this is a continuing war and not a one-time battle, our understanding of these microbes and the nature of disease will likely keep infectious disease at bay for the foreseeable future. Table 1.5 lists important events in the treatment and prevention of disease.
Table 1.5. Treatment and prevention of disease
| Year | Event |
| 1100 | Physicians in India and China realize that the liquid from the pustules of a smallpox victim, when scratched on the skin of a healthy patient, would most often cause mild disease. This intentional infection, termed variolation, would also give life-long protection against the illness. |
| 1721 | Lady Mary Wortley Montgue, wife of the ambassador to the Ottoman Empire, introduces variolation to Europe. |
| 1796 | Edward Jenner uses cowpox to immunize against smallpox. |
| 1884 | Ilya Ilich Metchnikoff demonstrates that certain body cells move to damaged areas of the body where they consume bacteria and other foreign particles. He calls the process phagocytosis. This is the beginning of the science of immunology, the study of the immune system. |
| 1885 | Paul Ehrlich proposes that certain chemicals affect bacterial cells and begins a search for one that can treat syphilis. |
| 1886 | Theobald Smith and D. E. Salmon develop a treatment for hog cholera by injecting killed hog cholera microorganisms into pigeons and demonstrate immunity to subsequent administration of a live microbial culture of cholera. |
| 1891 | Ehrlich shows that antibodies are responsible for part of immunity. |
| 1897 | Almwroth Wright and David Sample develop an effective vaccine against typhoid fever using killed cells of Salmonella typhi. |
| 1897 | Waldemar Haffkine develops a vaccine against the plague. |
| 1912 | Paul Ehrlich announces the discovery of a cure for syphilis. The cure is the first specific chemotherapeutic agent for a bacterial disease. |
| 1929 | Alexander Fleming publishes the first paper describing penicillin. |
| 1935 | Gerhard J. Domagk uses Prontosil, a chemically synthesized antimetabolite, to kill Streptococcus in mice. |
| 1938 | Max Theiler produces a vaccine against yellow fever by passaging the virus through mice to weaken it. |
| 1940 | Howard Florey and Ernest Chain produce an extract of penicillin and show it can kill bacteria in animals. |
| 1940 | Ernest Chain and E.P. Abraham describe a substance from E. coli that can inactivate penicillin. This demonstrates how rapidly bacteria can become resistant to antibiotics. |
| 1940 | Selman Waksman and H. Boyd Woodruff discover actinomycin, the first antibiotic obtained pure from a group of soil organisms, the actinomycetes. In subsequent years many antibiotics are isolated from this group including tetracycline and streptomycin. |
| 1941 | Charles Fletcher demonstrates that penicillin is non-toxic to human volunteers, by injecting a police officer suffering from a lethal infection. |
| 1942 | Selman Waksman suggests the word "antibiotic" to describe the class of compounds produced by one microorganism that inhibit or kill other microorganisms. |
| 1944 | Albert Schatz, E. Bugie, and Selman Waksman discover streptomycin, a very effective drug against tuberculosis. |
| 1944 | W. H. Feldman and H. C. Hinshaw at the Mayo Clinic successfully treat tuberculosis with streptomycin. |
| 1957 | The Soviet delegation to the World Health Organization proposes a vaccination effort to eradicate smallpox. The program finally begins in 1967. |
| 1977 | Ali Maow Maalin, age 23 of Somalia, is the last known victim of naturally occurring smallpox. |
| 1979 | Smallpox is declared to be eliminated. This is the only example of a microbial disease that has been wiped from the face of the Earth. (However, the recent specter of bioterrorism and the smallpox stocks kept by several governments make new epidemics of smallpox still possible.) |
Key Takeaways
- Variolation, immunization against smallpox, was a common practice before vaccination was common. This worked because the patient was exposed to a weak strain of smallpox, which did not kill, yet provided immunity to the disease.
- Edward Jenner discovered that cowpox could protect against smallpox, with a much lower incidence of complications than variolation.
- Pasteur discovered a general method for immunizing people against disease while working on chicken cholera. He coined the term vaccination to describe the technique.
- Paul Ehrlich spent 17 years in search of a chemical treatment against syphilis, eventually discovering salvarsan.
- Alexander Fleming discovered the first antibiotic, penicillin.
1 - 9 Environmental Microbiology
Learning Objectives
After reading this section, students will be able to...
- Describe the contributions of Martinus Beijerinck and Sergei Winogradsky to environmental microbiology.
- Explain why Linus Pauling and Emile Zuckerkandl thought macromolecules were a good way to compare and classify microbes.
- Describe how Carl Woese and others built on this work and developed a classification system based upon 16S rRNA.
Beijerinck and Winogradsky initiate the field of environmental microbiology
Contrary to what you would believe from news and advertising, the vast majority of bacteria are harmless to humans, and a large number are helpful. Understandably, early work in microbiology focused on pathogenic microbes and treatment of the diseases they caused. Martinus Beijerinck and Sergei Winogradsky began the transition from this early work into the present era, where we study the molecular biology of a wide variety of microorganisms. Both were interested in microbes present in the soil and water, and their work founded the discipline of environmental microbiology. A great deal of what we understand about bacteria in the environment and their impact can be traced back to the efforts of these two scientists.
Martinus Beijerinck, depicted in Figure 1.17, was originally trained as a botanist and began studying the microorganisms present in and around plants. He soon began experiments with microbes in the soil. His greatest contribution was the development of enrichment media. Previously, microbes were cultivated on media consisting of potatoes or extracts of leftover animal renderings. Such media would support the growth of many different bacteria, with chance and population density dictating what became dominant in the culture. In many cases, this was desirable, but in other cases, it was of interest to find bacteria capable of performing certain chemical conversions.

Figure 1.17. Martinus Beijerinck. The Dutch microbiologist Martinus Beijerinck. Drawing by Tammi Henke.
Beijerinck discovered that by adding or removing certain compounds from the medium or incubating under different conditions, it was possible to favor the growth of certain microbes and prevent the growth of others. An example is Beijerinck's work with nitrogen-fixing bacteria, which are important in agriculture and the global cycling of nitrogen. They are capable of taking nitrogen gas (N2) from the air and reducing it to ammonia (NH3). Nitrogen fixation is important since most organisms, including agriculturally important plants, can only use reduced nitrogen compounds such as ammonia. Beijerinck wanted to isolate an organism capable of fixing nitrogen in the presence of air because all previous isolates fixed nitrogen only under anaerobic conditions. By making a medium that did not contain a source of fixed nitrogen and then incubating in the presence of air (containing N2), he demanded that any microbe growing in the medium had to be able to derive its nitrogen by performing aerobic nitrogen fixation. Using this method, he succeeded in isolating a new microbe (Azotobacter chroococum) with these capabilities. Similar selective culture techniques (as he liked to call them) enabled his laboratory to isolate sulfur-reducing and sulfur-oxidizing bacteria, Lactobacillus species, green algae and many other microbes. Selective culture techniques have since been applied to many different groups of microorganisms, allowing them to easily be brought into pure culture.
Sergei Winogradsky, pictured in Figure 1.18, was also interested in soil bacteria, especially those involved in cycling nitrogen and sulfur compounds. He was one of the first to isolate microorganisms responsible for converting these elements in the soil, obtaining pure cultures of bacteria capable of oxidizing ammonia to nitrate. Winogradsky also studied the consumption of hydrogen sulfide gas by sulfur-oxidizing bacteria directly in their natural habitat.

Figure 1.18. Sergei Winogradsky. The Russian microbiologist Sergei Winogradsky. Drawing by Tammi Henke.
When working with Beggiatoa (one of these sulfur-oxidizing bacteria) he discovered it was obtaining its cell carbon from carbon dioxide. He used the term autotrophy to describe this property, a radical idea at the time, and one not readily accepted by other microbiologists. This led Winogradsky to propose the concept of chemolithotrophy; bacteria capable of growth using purely inorganic sources of carbon and energy. He realized that these microbes were essential for the cycling of nutrients in the environment, and the living world was dependent upon their action, long before others accepted the idea.
As a result of the work of Winogradsky and Beijerinck, there was great enthusiasm for identifying and classifying the bacteria inhabiting our natural world. For the first part of the 20th century many scientists isolated microbes and made proposals for their organization into genera and species. In 1909 Sigurd Orla-Jensen suggested a classification scheme based on the functions and abilities of the bacteria, such as growth on certain compounds or the production of specific by-products. The Society of American Bacteriologists, later to become the American Society of Microbiology, applied this technique to prepare a report on the classification of bacteria. This report eventually evolved into Bergey's Manual of Determinative Bacteriology in 1923 and subsequent editions have become authoritative reference works.
There was great optimism that as microbiologists brought more bacteria into pure culture, a clear organization based on the evolutionary history of microbes would emerge from studies of their physiology. However, by the 1940's it became clear that bacteria were unwilling to go along with this idea because classification based on one set of properties was often inconsistent with classification using a different set of properties. Scientists threw up their hands and gave up the idea of finding a system of classification based on growth and morphological characteristics.

Figure 1.19. Carl Woese. The American microbiologist Carl Woese, who applied the concept of molecular phylogeny to 16S rRNA and pioneered modern microbial taxonomy. (Source: University of Illinois-Champaign-Urbana)
Several decades passed during which bacteria became tools for understanding life, but their place in evolution was of little interest. Then in 1961 Brian McCarthy and E. T. Bolton developed methods for comparing genetic material between species, and Linus Pauling and Emile Zuckerkandl formalized the idea of using the makeup of biological molecules, their sequence, as a way to determine phylogenetic relationships. Later, Carl Woese, pictured in Figure 1.19, began to use this insight, choosing the sequence of ribosomal RNA as the basis for evolutionary comparisons. His initial efforts used the smaller RNA, termed 16S, of bacterial ribosomes, which he showed to be a marvelous "evolutionary clock." This analysis was a watershed event for the evolutionary classification of bacteria. In 1977 Woese used this technique to suggest dividing known bacteria into two separate domains that we now call Bacteria and Archaea, radically changing our view of the microbial world. Because all organisms have ribosomal RNA, scientists now apply this method of analysis to the identification and classification of large classes of organisms such as protozoa, fungi, plants and birds, fundamentally reshaping our views of the biological world. In subsequent years determination of 16S rRNA sequences, and then complete genomes, of microorganisms has enabled their natural classification. A review by Olsen, Woese, and Overbeek provided a catalyzing summary of the tree of life for microbes, including eukaryotic microorganisms. This reorganization has reshaped our view of the tree of life for all organisms and helped reassert the important role of microbes in shaping the past and affecting the earth's future. A renewed interest in environmental microbiology began in the 1980's and molecular techniques developed in the ensuing decades have allowed scientists to finally get a true picture of the environment. Landmark studies by Norman Pace, J. Craig Venter, and others have revealed the dominance of microbes in the environment. Many isolates are unculturable, and many of the proteins that are part of these microbes perform unknown functions. We have much to learn about our environment. Table 1.6 lists the significant events in environmental microbiology.
Table 1.6 Significant Events in Environmental Microbiology.
| Year | Event |
| 1887 | Sergei Winogradsky studies Beggiatoa and establishes the concept of autotrophy. |
| 1888 | Martinus Beijerinck develops the technique of enrichment culture. |
| 1891 | Winogradsky discovers the organisms responsible for nitrification is soil, which is of great importance in agriculture because nitrogen is a limiting nutrient in the soil. |
| 1904 | Martinus Beijerinck obtains the first pure culture of sulfur-oxidizing bacterium, Thiobacillus denitrificans. |
| 1904 | Cornelius Johan Koning suggests that fungi are critical for the decomposition of organic matter. |
| 1909 | Sigurd Orla-Jensen proposes the use of physiological characteristics for the classification of bacteria. He later publishes a monograph on lactic acid bacteria that establishes the criteria for assignment. |
| 1920 | The Society of American Bacteriologists presents a report on the characterization and classification of bacterial types that becomes the basis for Bergey's manual in 1923. |
| 1961 | Brian McCarthy and E. T. Bolton describe a method to compare genetic material from different species using hybridization. Using this technique it is possible to quantitatively compare the relatedness of the two species. |
| 1965 | Emile Zuckerkandl and Linus Pauling publish "Molecules as documents of evolutionary history", making a compelling case for the use of molecular sequences of biological molecules to determine evolutionary relationships. |
| 1969 | Don Brenner and colleagues establish a more reliable basis for the classification of clinical isolates among members of the Enterobacteriaceae. They use nucleic acid reassociation, where DNA of one organism is allowed to hybridize with another organism. This technique is used to help define a species. |
| 1977 | Carl Woese uses ribosomal RNA analysis to identify a third form of life, the Archaea, whose genetic makeup is distinct from but related to both Bacteria and Eukarya. |
| 1977 | Holger Jannasch discovers abundant life at the bottom of the ocean near deep sea hydrothermal vents. The entire system is dependent upon sulfur oxidizing microorganisms. Light and photosynthesis do not drive the process. |
| 1982 | Karl Stetter isolates hydrothermophilic microbes (Archaea) that can grow at 105°C. The discovery redefines the upper temperature at which life can exist. |
| 1994 | Gary Olsen, Carl Woese and Ross Overbeek summarize the state of phylogeny in prokaryotes. This causes scientists to rethink the classification of life and emphasizes the importance of microbes. |
Key takeaways
- Martinus Beijerinck and Sergei Winogradsky were two of the major scientists investigating microbes in the environment that did not cause disease.
- Linus Pauling and Emile Zuckerkandl developed the idea of using the sequence of biological macromolecules to compare and classify microbes.
- Carl Woese and others then developed a classification system based upon 16S rRNA. This system organized the tree of life and is used today to help classify all organisms.
1 - 10 Molecular Microbiology
Learning Objectives
After reading this section, students will be able to...
- Explain how microbes have been critical tools for the understanding of basic biology at the molecular level.
- Describe Fred Griffith transformation experiments and their significance..
- Explain the the one gene-one enzyme hypothesis of Beadle and Tatum
- Explain why E. coli was, and continues to be, a useful model microorganism for understanding basic questions about biology.
- Describe the discoveries of Joshua Lederberg.
- Describe the development of molecular biology and give examples of its uses.
- Describe the development of DNA sequencing and give examples of its uses.
In the early part of the 20thcentury techniques were developed to examine the inner workings of the cell and researchers performed much of the work in bacteria due to their experimental accessibility. Before this period, the method of turning genetic information into the proteins that carry out cellular processes was completely unknown. Indeed, the chemical composition of the genetic material was being hotly debated. From Gregor Mendel's work with pea plants, we understood the nature of inheritance and heredity. However, it was unclear what molecule in the cell maintained and passed hereditary information on to subsequent generations. The leading contender for this role was protein, when in 1928 Fred Griffith discovered transformation in bacteria.
Griffith knew that when Streptococcus pneumoniae strains are injected into mice, they cause rapid deterioration and death. Griffith isolated strains of the bacteria that were no longer capable of producing an outer slime layer and appeared rough when grown on solid medium in contrast to the smooth colonies of the original isolate. When he injected the rough strains into mice, no illness resulted. Similarly, when a smooth microbe was heat-killed and then injected, the mice showed no signs of infection. A surprise was awaiting Griffith when he injected a mixture of dead cells of a smooth isolate with live cells of a rough isolate into mice: they died. When bacteria were isolated from these dead mice, they formed smooth colonies. Griffith hypothesized that the ability to create the slime layer was passed from the dead smooth cells to the viable rough cells, making them pathogenic again. The process became known as transformation.
Griffith was ridiculed, as most scientists believed his preparation was contaminated with viable smooth cells. It was not until 1944 that his student Oswald Avery and coworkers repeated the experiments of Griffith, reproducing his results and discovering that DNA was the material from the dead smooth cells that transformed the rough mutant. This transformation was strong proof that the hereditary material in cells was DNA. Further experiments in the 1940's clearly established this observation with Joshua Lederberg discovering two more ways that DNA could be transferred between bacterial cells, conjugation and transduction. For more information on these experiments, see the chapter on Genomics and Genetics
In 1943, Beadle and Tatum reported experiments with the fungus Neuropsora crassa that eventually established the idea that each gene in the DNA typically codes for one protein (the one gene-one enzyme hypothesis). About ten years later, Francis Crick, Rosalind Franklin, James Watson, and Maurice Wilkins worked on experiments describing the structure of DNA and hypothesized about its replication. It was now clear that DNA stored the information for proteins and that proteins performed the many functions of the cell.
The important question now became, how does one convert the information in DNA into protein. A significant contribution in understanding this puzzle were the cell-free systems developed by Paul Zemecnik and his laboratory. The cell-free systems first used rat liver and later the bacterium E. coli. He and other scientists used these systems to study translation in the test tube, discovering many of the important molecules involved in the process. An intense effort to describe these molecular events then ensued. The first insight came when Crick, Sydney Brenner, and colleagues proposed the existence of transfer RNA (tRNA). This molecule helps to create amino acid polymers based on the nucleic acid sequence. Brenner, Francois Jacob, and Matthew Meselson revealed another important part of the puzzle when they discovered that the ribosome was the site of translation and that the molecule it translated is RNA, not DNA. Marshall Nirenberg and J. H. Matthaei solved the next mystery by when they developed methods to decipher the genetic code that dictates the correspondence of nucleic acids to amino acids. After the stunningly productive decade of the 1960's, we now understood the framework for converting genetic information into proteins. Further work demonstrated that this basic mechanism is conserved across all biology. For more information on the process of transcription and translation, see the chapter on the Central Dogma
Table 1.7 lists important events in learning about the central molecular events in biology. Note that almost all of this work occurred in microorganisms.
Table 1.7 Significant Events in Learning about the Molecules of Life
| Year | Event |
| 1928 | Frederick Griffith discovers transformation in bacteria and establishes the foundation of molecular genetics. |
| 1941 | George Beadle and Edward Tatum develop the one-gene one-enzyme hypothesis. |
| 1943 | Salvador Luria and Max Delbruck demonstrate that inheritance in bacteria follows Darwinian principles. |
| 1944 | Oswald Avery, Colin MacLeod, and Maclyn McCarty show that DNA is the hereditary material. |
| 1946 | Joshua Lederberg and Edward L. Tatum discover a second method of gene transfer in bacteria: conjugation. |
| 1952 | Joshua Lederberg and Norton Zinder find a third method of DNA transfer in bacteria using bacteriophage: transduction. |
| 1952 | Alfred Hershey and Martha Chase suggest that only DNA is needed for viral replication. |
| 1953 | Francis Crick, Maurice Wilkins and James Watson describe the structure of DNA. |
| 1954 | Paul Zemecnik develops a cell-free system for translation using rat liver |
| 1954 | Francis Crick, Sydney Brenner, and colleagues propose the existence of RNA that helps to transfer the information in DNA into protein (tRNA). |
| 1956 | Paul Zemecnik discovers tRNA in his cell-free system |
| 1957 | Francois Jacob and Elie Wollman show that the chromosome of E. coli is circular. |
| 1959 | Peter Mitchell proposes the chemiosmotic theory in which a molecular process is coupled to the transport of protons across a biological membrane. He argues that this principle explains ATP synthesis, solute accumulations or expulsions, and cell movement. |
| 1959 | Arthur Pardee, Francois Jacob, and Jacques Monod show that lactose induces β-D-galactosidase the catabolic enzyme that begins the degradation of the sugar. |
| 1960 | Arthur Kornberg demonstrates DNA synthesis in cell-free bacterial extracts and later shows a specific enzyme complex catalyzes the synthesis of DNA. |
| 1960 | Francois Jacob, David Perrin, Carmen Sanchez and Jacques Monod propose a mechanism for the for control of bacterial gene expression in an organization they call the operon. |
| 1960 | Paul Zemecnik and Robert Lamberg develop a bacterial cell-free system using E. coli |
| 1961 | Marshall Nirenberg and J.H. Matthaei observe that a synthetic polynucleotide, composed only of a string of the base uracil, directs the synthesis of a polypeptide composed only of phenylalanine. This begins the quest to unravel the genetic code that translates DNA into protein. |
| 1961 | Sydney Brenner, Francois Jacob and Matthew Meselson show that ribosomes are the site of protein synthesis and that RNA carries messages from the DNA to the ribosome. |
| 1968 | Lynn Margulis proposes that endosymbiosis has led to the generation of mitochondria and chloroplasts from bacterial progenitors. |
| 1970 | Howard Temin and David Baltimore independently discover reverse transcriptase, a radically different way to alter genetic information in cells. |
| 1975 | Thomas Cech and Sidney Altman independently show that RNA, and not just protein, can serve directly as a reaction catalyst. |
Work on microbes paves the way to taking control of the genome
One final thread we will trace is the rise of molecular microbiology and the technique of genomics. This pathway begins in 1885 with the seemingly unimportant isolation of a common intestinal microorganism by Theodor Escherich. The microbe is eventually renamed Escherichia coli in his honor. Little did Dr. Escherich know at that time, but this bacterium would play a central role in our understanding of life. E. coli achieved this position because it has several properties that made it invaluable as an experimental system:
- E. coli is easy to cultivate. It can grow on a wide variety of media, at temperatures ranging from 20-45 °C, in the presence or absence of air.
- E. coli grows quickly. Under normal laboratory conditions, it goes through a complete life cycle and doubles its population about every 30 minutes. With this rapid growth rate, it is possible to generate large populations in just a few hours. Researchers can set up most experiments one day and observe results the next.
- It can survive storage for long periods of time without losing viability using relatively simple preservation techniques.
- It was available. There is an element of arbitrariness that this microbe came to serve such a vital role in biology. Many other bacteria probably could have filled this position, but E. coli was the one chosen.
Several key discoveries laid the foundation for future developments in molecular biology. The first of these was the discovery of autonomously replicated pieces of DNA separate from the bacterial chromosome. Plasmids, as Joshua Lederberg called them, were capable of carrying genes that could be regulated and moved from one microbe to another independently of the bacterial chromosome. The ability of some plasmids to move between different cells meant that different bacteria could rapidly inherit genes carried on these plasmids. Scientists later found that one class of plasmids encoded resistance genes that would enable bacteria to grow in the presence of an antibiotic. It later became clear it was possible to select for the presence of these plasmids by demanding growth of the bacterial strain on a medium containing the antibiotic to which the plasmid encoded resistance.
A second discovery that greatly impacted modern genetic engineering was that bacteria have their own parasites. As mentioned earlier, some viruses infect and kill bacteria, just as viruses infect animals and plants. The analysis of bacterial viruses was a major avenue to understanding the bacteria themselves.
For example, investigation of the ability of some viruses to grow well on certain strains of bacteria and not others lead to the discovery of restriction-modification systems in bacteria. These systems allow bacteria to recognize and destroy foreign DNA and consist of two parts. A modification enzyme that labels DNA as belonging to the bacterium and a restriction enzyme that recognizes a certain 4-8 base pair sequence in the DNA and cuts it in two if it has not been modified. The landmark paper of Hamilton-Smith and Kent Wilcox showed that restriction enzymes could cleave double-stranded DNA into discrete pieces. The authors correctly hypothesized that the enzyme was recognizing a sequence of DNA and cutting it.
Further studies on restriction enzymes led to the birth of genetic engineering. Janet Mertz and Ronald Davis made restriction enzymes into tools for the manipulation of DNA when they showed that many of the breaks these enzymes made in the DNA produced single-stranded complementary ends. Stanley Cohen, Annie Chang, Robert Helling, and Herbert Boyer then demonstrated that restriction enzymes could break any DNA into fragments, and by mixing that with a plasmid digested similarly, it was possible to create recombinant molecules. Moving the plasmid into a microbe and growing it on a selective medium could produce any desired amount of these recombinant molecules.
Another discovery was "brewing" in the western United States. In the mid-1960s, Thomas Brock was on a trip to Yellowstone and became intrigued by the mats of green, brown, and pink material that he found in many of the hot springs. Brock was sure that these were living communities of microorganisms, yet the springs were at temperatures near boiling. Subsequent research proved his hypothesis correct and led to the isolation of a microbe, Thermus aquaticus, capable of growth at 85 °C.
Organisms capable of growing at this high temperature were unheard of at the time, and the discovery spawned two major developments. First, work with T. aquaticus and other unusual microbes by Woese and many others lead to the discovery of the Archaea. Second, the enzyme responsible for replicating DNA in T. aquaticus is extremely heat stable, surviving temperatures of 95 °C and copying DNA at 72 °C. This enzyme allowed the next important breakthrough. In the early '70s, H. Gobind Khorana and colleagues had suggested a method to amplify DNA in a test tube. As they envisioned it, the method used short oligonucleotides, DNA polymerase, and repeated heating and cooling cycles. However, the cycle denatures the polymerase during each heating step so that the procedure would require massive amounts of prohibitively expensive enzymes. When Kary Mullis reinvented the method in 1985, the thermostable polymerase of T. aquaticus was readily available, which made the procedure wildly easier and cheaper to apply. He termed it polymerase chain reaction (always referred to as PCR), and it has become a pivotal technique in many areas of science, from detecting bacteria in the environment to the analysis of evidence at crime scenes.
In 1977, Walter Gilbert and Fred Sanger independently developed methods for determining the exact sequence of bases in DNA. These techniques became immediately useful in determining the sequence of numerous important genes being investigated in countless laboratories.
Further refinement has made DNA sequencing more efficient. It became efficient enough that in 1985 Robert Sinsheimer convened a meeting of several biologists active in genetics and gene mapping. Out of this meeting came a proposal to sequence the entire three billion base pairs of the human genome. This project was equivalent at the time of proposing a mission to the moon, technically possible but with an unpredictable value. Scientists estimated the project's initial cost to be $10 a base pair or a staggering 30 billion dollars. There was resistance from a significant portion of the scientific community, fearing that the project would siphon funds away from other worthy scientific projects. However, the clarity of the goal ignited the imagination of Congress and the public, creating unstoppable momentum for the project. After a decade of effort, a preliminary draft of the entire genome was released in the spring of 2001, earlier than projected and dramatically under budget. The project itself drove the development of sequencing technology, allowing the determination of millions of base pairs of sequence. Due to advances in DNA sequencing technology, costs have dropped to far below a fraction of a penny per base at large sequencing facilities.
Due to the ease of DNA sequencing, there are now over 100,000 sequenced genomes. These include the genomes of many pathogenic bacteria, the mouse, the fruit fly, and the nematode Caenorhabditis elegans, the last being an experimental model for development in eukaryotes. The use of sequence data to investigate biological problems has resulted in the development of the fields of genomics and bioinformatics. In their infancy, these fields are already producing astonishing findings that accelerate the progress of evolutionary biology, immunology, bacterial pathology, bacterial physiology, and cancer treatment. Table 1.8 lists important events in understanding the genome.
Table 1.8. Events in understanding genomes
|
Year |
Event |
| 1885 | Theodor Escherich isolates a microbe from the colon that is later given the name Escherichia coli in his honor. This microbe later becomes the workhorse of molecular biology. |
| 1897 | Edward Buchner helps launch the field of enzymology by developing a cell extract from yeast that is able to ferment sugar to alcohol. |
| 1952 | Salvador Luria and Mary Human, and independently Jean Weigle, describe sensitivity in bacteriophage imposed by the host on which it was grown. The viruses are restricted to grow well only on specific strains of bacteria. This later leads to the study of bacterial systems of restriction and modification, and eventually the discovery of restriction enzymes. |
| 1959 | O. Sawada and others demonstrate that antibiotic resistance can be transferred between Shigella strains and Escherichia coli strains by plasmids. |
| 1966 | Jon Beckwith and Ethan Signer move the lac region of E. coli into another microorganism to demonstrate genetic control. It is quickly realized that chromosomes could be redesigned and genes moved. |
| 1967 | Waclaw Szybalski and William Summers develop the technique of DNA-RNA hybridization (mixing nucleic acids together and allowing them to base pair) to investigate the bacteriophage T7. This technique finds wide use in many experiments. |
| 1967 | Thomas Brock identifies Thermus aquaticus, a bacterium that grows at 85 °C. Heat-stable DNA polymerase is later isolated and used in PCR. Investigation of this organism also leads to the discovery of the domain Archaea. |
| 1967 | Werner Arber shows that bacterial cells have enzymes capable of modifying DNA by adding methyl groups at cytosines and adenosines. This methylation helps the cell identify its own DNA. Accompanying nucleases recognize these sites and cut the DNA if it is not methylated. |
| 1970 | Hamilton Smith and Kent W. Wilcox describe the action of restriction enzymes, discovered by Arber, by the purification of one of these enzymes from Haemophilus influenzae. |
| 1972 | Janet Mertz and Ronald W. Davis establish that the RI restriction enzyme from Escherichia coli cuts at a specific site on the DNA. They also reveal that the cleaved ends of the DNA are complementary, opening the way for cloning. |
| 1972 | Paul Berg creates the first recombinant DNA molecule from viral and bacterial DNA. |
| 1973 | Stanley Cohen, Annie Chang, Robert Helling, and Herbert Boyer develop the process of gene cloning. |
| 1975-1976 | The Asilomar Conference is convened to discuss possible problems associated with gene cloning. A one-year moratorium is suggested, as well as guidelines for cloning research and for genetic engineering. |
| 1977 | Walter Gilbert and Fred Sanger independently develop methods for determining the sequence of DNA. |
| 1980 | The U. S. Supreme Court rules that microorganisms altered in the laboratory can be patented. |
| 1982 | U. S. Pharmaceutical manufacturer Eli Lilly markets the first genetically-engineered human insulin. |
| 1983 | Jeff Schell and Marc Van Montagu, Mary-Dell Chilton and colleagues move genes into plants. |
| 1988 | Kary Mullis uses a heat-stable enzyme from Thermus aquaticus to establish PCR technology. |
| 1992 | The entire sequence of one of the sixteen chromosomes of the yeast Saccharomyces cerevisiae is determined. |
| 1995 | Craig Venter, Hamilton Smith, Claire Fraser, and colleagues at TIGR elucidate the first complete genome sequence of a microorganism, Haemophilus influenzae. In the ensuing years many laboratories have produced sequences for dozens of microbes including many important pathogens and those of industrial or environmental importance. |
| 2000 | The human genome project begun in 1990 finished a working draft of the entire human genome. |
Let'send this chapter with a defense of fundamental scientific research. That is, research without an obvious applied goal or application. If you look at the initial results generated in many of these important analyses, the research would have seemed esoteric and inconsequential. Is the analysis of microbes in the colon important? Who cares why a virus can grow in one strain of bacteria and not another? Why should the government support Tom Brock's isolation of microbes from the hot springs of Yellowstone National Park? What use is knowing the DNA sequence of a virus or a bacterial operon? Yet, each of these insights began a thread of inquiry that changed the world. The take-home message is that basic research creates insights and applications beyond the imagination of even the scientists performing the work and should be supported.
Key Takeaways
- Microbes have been critical tools for the understanding of basic biology at the molecular level.
- Fred Griffith performed transformation experiments that suggested that DNA was the hereditary material. His student, Avery Oswald, later demonstrated this conclusively.
- Beadle and Tatum developed the one gene-one enzyme hypothesis using Neurospora crassa as the test microbe.
- Many scientists working with E. coli deciphered the nature of DNA, RNA and protein. These experiments lead to our understanding of how information in the cell is stored and how it is converted into proteins. E. coli continues to help scientists answer many basic questions about biology and is an indispensable tool in genetics and genomics.
- Joshua Lederberg discovered plasmids, small, circular, self-replicating pieces of DNA. The discovery of restriction enzymes allowed scientists to build chimeric DNA molecules and gave birth to the technique of cloning. It was now possible to isolate any DNA molecule, mix it with a plasmid and created recombinant molecules. These could be grown in E.coli, and other microbes, to make as many copies as the scientist needed.
- The polymerase chain reaction allows the amplification of any DNA sequence over a millionfold. PCR has found hundreds of applications in science and medicine.
- DNA sequencing, independently developed by Walter Gilbert and Fred Sanger, has allowed the determination of the sequence of millions of base pairs of DNA. Advances in sequencing technology now make it routine to sequence billions of base pairs in a single day. Genomic sequences of organisms, including humans and many important pathogens, are generating new insights into us and the diseases caused by certain microbes. This large amount of data has spawned the fields of genomics and bioinformatics.
1 - 11 Summary
Microbiology is the study of organisms that at some point in their life exist as single cells and contain a nucleic acid genome that can replicate. Many organisms fall into this definition including algae, fungi, protozoa, bacteria and archaea. Together these organisms have a profound impact on the biosphere, making up the majority of life both in number and total mass. Microbes cause many illnesses, and understanding these infections has led to cures and better treatments. The emergence of new infectious agents will spur continued interest in microbiology. Many more microbes grow harmlessly in the environment, taking advantage of chemicals and/or sunlight to grow. Research into these microbes has also helped us understand the basic framework of life and revealed the basic fundamental rules that govern living systems. In the past, microbes have been used in experiments to answer many scientific questions, and they will continue to serve as excellent tools of inquiry in the future. A significant number of these discoveries have led to important applications in many areas of human endeavor.
Microbiology is a fluid science
Microbiology is constantly changing. Most of what we will discuss consists of well-established principles, and it would be astounding if these changed. However, we have tried to keep this book on the cutting edge of the science of small things as possible. When you live this close to the edge, you are bound to get some cuts, and some of what you read here today, could change when more is known later. Keep that in mind.
If you enjoyed this free chapter on the the relevance and history of microbiology, consider buying the book, for only $30.